- Protocols
- Articles and Issues
- About
- Become a Reviewer
Current Issue in 2026
Volume: 16, Issue: 10
Biochemistry
PEPTERGENT: A Peptide-Based Reagent for Detergent-Free Extraction of Membrane Proteins and Purification of Membrane Proteomes
Bioinformatics and Computational Biology
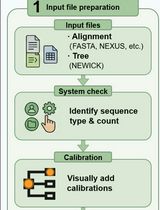
A Step-by-Step GUI-Based Protocol for Molecular Dating Analysis Using PhyloSuite v2
Biophysics
Using Single-Particle Fluorescence Microscopy to Quantify Substrate Binding of Peptidoglycan-Modification Enzymes
Cell Biology
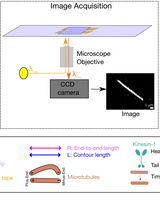
Quantification of Spatial Patterns of Microtubule Transport by Kinesin-1 Head and Tail
Isolation and Biophysical Characterization of Extracellular Vesicles Released by Myocytes
Immunology
A Versatile In Vitro Quantitative Assay for Macrophage Efferocytosis in Diverse Research Applications
Microbiology
An In Vitro A-431 Epithelial Cell Infection Model for Studying Fungal Pathogenicity and Immune Responses Associated With Vulvovaginal Candidiasis
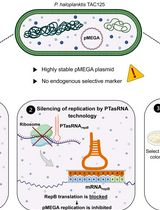
Plasmid Curing of Pseudoalteromonas haloplanktis TAC125 Using Homologous Recombination and PTasRNA Gene Silencing
Neuroscience
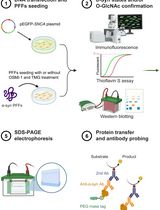
Chemoenzymatic Labeling Method for Detection of O-GlcNAcylated α-Synuclein Proteins by Western Blot
Plant Science
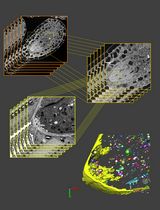
3D Reconstruction of Mature Arabidopsis Ovules Using FIB-SEM to Study Filiform Apparatus Morphology
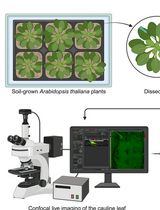
Analysis of Cauline Leaf Development in Arabidopsis thaliana Using Time-Lapse Confocal Microscopy
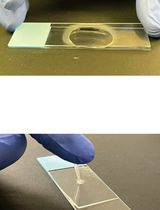
Biolayer Interferometry (BLI) to Quantify RALF1–Pectin Interactions
Correction