- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
A Rapid and Visual Soybean Hairy Root Transformation Protocol Using the RUBY Reporter
Published: Vol 16, Iss 6, Mar 20, 2026 DOI: 10.21769/BioProtoc.5635 Views: 10
Reviewed by: Ying LiAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Agrobacterium-mediated Transformation of Japonica Rice Using Mature Embryos and Regenerated Transgenic Plants
Ammar Elakhdar [...] Takahiko Kubo
Sep 20, 2021 6249 Views

Agrobacterium-mediated Genetic Transformation of Cotton and Regeneration via Somatic Embryogenesis
Alka Srivastava [...] Praveen C. Verma
May 20, 2023 4390 Views
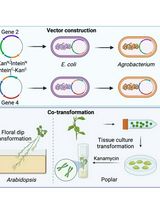
A Novel Gene Stacking Method in Plant Transformation Utilizing Split Selectable Markers
Guoliang Yuan [...] Xiaohan Yang
Feb 20, 2025 1994 Views
Abstract
Agrobacterium rhizogenes–mediated hairy root transformation provides a rapid platform for gene function analysis prior to stable whole-plant transformation. However, most existing hairy root transformation methods rely on tissue culture and require chemical or fluorescence-based selection, which increases experimental complexity. Here, we describe a tissue culture–free soybean hairy root transformation protocol incorporating the RUBY visual reporter system. While this work does not introduce a new transformation concept, it presents a streamlined implementation of established soybean hairy root methodologies that emphasizes procedural simplicity, reduced handling, and faster access to functional root material. Transgenic roots expressing RUBY can be directly identified by red pigmentation with the naked eye. In RUBY-positive roots, candidate genes driven by the CaMV 35S promoter showed higher expression levels than those in empty-vector controls, indicating that the system supports effective gene expression. Using this procedure, clearly identifiable transgenic hairy roots can be obtained within 20 days. Overall, this protocol simplifies induction and screening while reducing operational complexity and equipment requirements.
Key features
• Induces soybean hairy roots without tissue culture, simplifying the transformation workflow.
• Incorporates RUBY reporter for direct naked-eye identification of transgenic roots.
• Generates identifiable transgenic hairy roots within 20 days, enabling rapid functional analysis.
• Applicable to gene and protein function studies, metabolite profiling, and root–microbe interaction assays.
Keywords: Agrobacterium rhizogenesGraphical overview
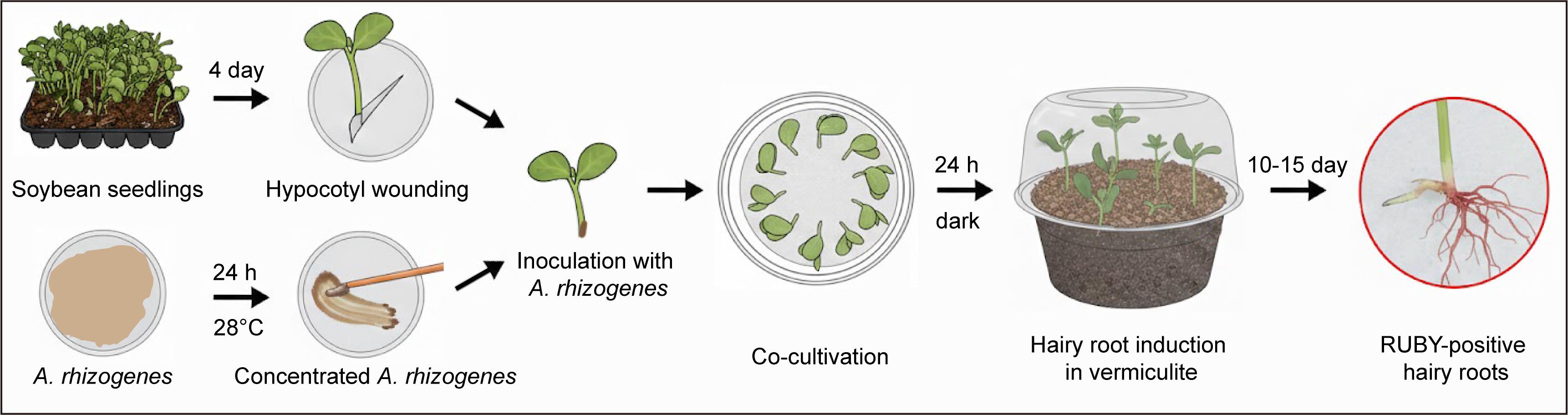
Schematic diagram illustrating the workflow of soybean hairy root transformation using the RUBY reporter
Background
Agrobacterium rhizogenes is a gram-negative soil bacterium belonging to the family Rhizobiaceae that infects plants through wound sites. During infection, transfer DNA (T-DNA) carried on the Ri plasmid is delivered into plant cells and randomly integrated into the host genome, where the introduced genes are expressed and induce the formation of roots with sustained growth capacity [1,2]. Because this process bypasses the requirement for whole-plant regeneration, A. rhizogenes–mediated hairy root transformation enables rapid generation of root tissues with stable transgene expression.
Hairy root transformation has therefore become a widely used experimental system in plant functional genomics. The approach is applicable to diverse molecular and physiological studies, including protein expression, subcellular localization, and protein–protein interaction analyses [3,4], secondary metabolism pathway investigation [5,6], and root–microbe interaction studies [7,8]. In addition, hairy root systems have been increasingly applied in genome editing research as a convenient platform for evaluating CRISPR/Cas9 single-guide RNA (sgRNA) efficiency, with editing outcomes that show good consistency with stable transformation results [8,9].
In soybean [Glycine max (L.) Merr.], conventional whole-plant stable transformation is time-consuming, labor-intensive, genotype-dependent, and often exhibits low efficiency, which limits its application in rapid gene function analysis and high-throughput studies [10,11]. A. rhizogenes–mediated hairy root transformation overcomes many of these limitations, providing an efficient alternative for root-focused studies, including analyses of metabolism, abiotic stress responses, and plant–microbe interactions [12–16]. Nevertheless, many existing soybean hairy root protocols still involve tissue culture steps and require antibiotic selection or fluorescence microscopy to identify positive transformants, with overall experimental durations typically ranging from 30 to 45 days [17,18].
To address these limitations, we built upon a tissue culture–free soybean hairy root induction system previously reported by Chen et al. [19] and incorporated the RUBY visual reporter gene for direct identification of transgenic roots. RUBY is a synthetic reporter comprising three betalain biosynthetic enzymes, CYP76AD1, DODA, and glucosyltransferase, which together drive the accumulation of a vivid red pigment in transformed cells [20,21]. Because RUBY expression can be readily detected by the naked eye, it provides an effective indicator for transformation events and transgene expression levels. Incorporation of the RUBY reporter eliminates the need for antibiotic or fluorescence-based selection, simplifies experimental procedures, and reduces equipment requirements, thereby offering a streamlined and reproducible method for generating transgenic soybean hairy roots suitable for downstream functional analyses.
Materials and reagents
Biological materials
1. Soybean seeds (Williams 82)
2. Agrobacterium rhizogenes K599 (Coolaber Science & Technology, catalog number: CC410)
3. E. coli DH5α competent cell (Sangon Biotech, catalog number: B528413)
4. pCAMBIA1300 plasmid (CAMBIA, stored in our laboratory)
Reagents
1. Xho I restriction enzymes (New England Biolabs, catalog numbers: R0146); store at -20 °C
2. Kanamycin sulfate (Sangon Biotech, catalog number: A600286); store at 4 °C
3. Streptomycin sulfate (Sangon Biotech, catalog number: A610494); store at 4 °C
4. Tryptone (OXOID, catalog number: LP0042B); store at room temperature (RT)
5. Yeast extract (OXOID, catalog number: LP0021T); store at RT
6. Sodium chloride (NaCl) (Sangon Biotech, catalog number: A501218); store at RT
7. Calcium chloride dihydrate (CaCl2·2H2O) (Sangon Biotech, catalog number: A501330); store at RT
8. Glycerol (Sangon Biotech, catalog number: A600232); store at RT
9. Sodium hypochlorite (NaClO) (Sangon Biotech, catalog number: A501944); store at 4 °C
Caution: NaClO is a strong oxidizing agent.
10. Hydrochloric acid (HCl), 37% (Sigma-Aldrich, catalog number: 320331); store at RT
Caution: HCl is highly corrosive.
11. Acetosyringone (AS) (Beijing KEHBIO Technology, catalog number: K91068); store at 4 °C
12. Agar powder (Beijing Solarbio Science & Technology, catalog number: A8190); store at RT
13. KOD-Plus DNA polymerase (TOYOBO, catalog number: KOD-201); store at -20 °C
14. EZ-10 Spin Column DNA Gel Extraction kit (Sangon Biotech, catalog number: B610353); store at RT
15. Agarose (Sangon Biotech, catalog number: A620014); store at RT
16. ClonExpress MultiS One Step Cloning Kit (Vazyme Biotech, catalog number: C113); store at -20 °C
17. Dimethyl sulfoxide (DMSO) (Diamond, catalog number: A100231); store at RT
Caution: DMSO can facilitate dermal absorption of chemicals and may cause skin irritation.
18. Tris base (Sangon Biotech, catalog number: A501492); store at RT
19. Glacial acetic acid (Sinopharm Chemical Reagent, catalog number: 10000208); store at RT
20. EDTA disodium salt dihydrate (Sangon Biotech, catalog number: A610185); store at RT
Solutions
1. Liquid and solid lysogeny broth (LB) medium (see Recipes)
2. 0.1 M NaCl solution (see Recipes)
3. 20 mM CaCl2 solution (see Recipes)
4. AS stock solution (see Recipes)
5. Kanamycin stock solution (see Recipes)
6. Streptomycin stock solution (see Recipes)
7. 50× TAE buffer (see Recipes)
Recipes
1. Liquid and solid LB medium
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Tryptone | 10 g/L | 10 g |
| NaCl | 10 g/L | 10 g |
| Yeast extract | 5 g/L | 5 g |
| Agar powder (for solid LB only) | 15 g/L | 15 g |
| Deionized water | n/a | Up to 1L |
Note: Autoclave the medium at 121 °C for 15 min and store at RT for up to one month.
2. 0.1 M NaCl solution
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| NaCl | 0.1 M | 5.84 g |
| Deionized water | n/a | Up to 1 L |
Note: Filter-sterilize the solution using a 0.22 μm syringe filter and store at 4 °C for six months.
3. 20 mM CaCl2 solution
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| CaCl2·2H2O | 20 mM | 0.29 g |
| Deionized water | n/a | Up to 100 mL |
Note: Filter-sterilize the solution using a 0.22 μm syringe filter and store at 4 °C for six months.
4. AS stock solution
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| AS | 100 mg/mL | 1 g |
| DMSO | n/a | Up to 10 mL |
Note: Filter-sterilize the solution using a 0.22 μm syringe filter, aliquot into 500 μL portions, and store at -20 °C and protected from light for up to six months.
5. Kanamycin stock solution
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Kanamycin sulfate | 50 mg/mL | 2.5 g |
| Deionized water | n/a | Up to 50 mL |
Note: Filter-sterilize the solution using a 0.22 μm syringe filter, aliquot into 500 μL portions, and store at -20 °C for six months.
6. Streptomycin stock solution
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Streptomycin sulfate | 50 mg/mL | 2.5 g |
| Deionized water | n/a | Up to 50 mL |
Note: Filter-sterilize the solution using a 0.22 μm syringe filter, aliquot into 500 μL portions, and store at -20 °C for six months.
7. 50× TAE buffer
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Tris base | 2.0 M | 242 g |
| Glacial acetic acid | 1.0 M | 57.1 mL |
| EDTA (disodium salt, pH 8.0) | 50 mM | 100 mL of 0.5 M EDTA |
| Deionized water | n/a | Up to 1 L |
Note: Dilute 50× TAE buffer to 1× with deionized water before use. Store 50× stock at RT for six months.
Laboratory supplies
1. Glass dishes (12 cm dish) (WuYi)
2. Filter paper, 11 cm (NEWSTAR)
3. 1.5 mL microcentrifuge tubes (Axygen, catalog number: MCT-150-C)
4. 0.2 mL microcentrifuge tubes (Axygen, catalog number: PCR-02-C)
5. 50 mL microcentrifuge tubes (Biosharp, catalog number: CT-002-50A)
6. 0.22 μm syringe filters (Merck Millipore, catalog number: SLGPR33RB)
7. Plastic spreader (Biosharp, catalog number: BS-PS-A)
8. Sterile Petri dishes (9 cm dish) (Biosharp, catalog number: BS-90-D)
9. Surgical blades (Aladdin, catalog number: C3028-02-100EA)
10. Disposable plastic cups (CHEMATE, catalog number: 0861-1119)
11. Vermiculite (Santaibio, catalog number: ZS-02-04)
12. Nutrient soil (Scotts Miracle-Gro, universal type)
13. 250 mL Erlenmeyer flask (PYREX, catalog number: 4980-250)
Equipment
1. Microvolume spectrophotometer (IMPLEN, model: NanoPhotometer® N50)
2. Growth chamber (HilPoint, model: FH-600)
3. Centrifuge (HERMLE Labortechnik GmbH, model: Z216MK)
4. Temperature-controlled shaking incubator (XiangShang, model: XS-ZD3)
5. Biochemical incubator (BLabotery, model: SPL-80)
6. Ultra-low temperature freezer (Sonyo Refrigeration, model: MDF-682)
7. Autoclave (Yamato, model: SQ510C)
8. Gel imaging system (Shenhua Science Technology, model: SH-520)
9. PCR thermal cycler (Bio-Rad Laboratories, model: T100)
10. Nucleic acid electrophoresis system (Beijing Junyi-Dongfang Electrophoresis Equipment, model: JY600E)
11. Chemical fume hood (WuXi Jinhongyang Technology, model: BM220)
Procedure
A. Construction of pCAMBIA1300-RUBY expression vector
Note: In our work, the pCAMBIA1300 vector was used as the backbone, in which the hygromycin resistance (Hyg) gene was replaced by the RUBY reporter cassette. RUBY expression was driven by the cauliflower mosaic virus 35S (CaMV 35S) promoter and terminated by the CaMV polyadenylation signal (Figure 1A). The detailed construction procedure is described below.
1. Linearize the pCAMBIA1300 backbone: Digest the pCAMBIA1300 vector backbone by combining 20 μL of plasmid DNA (500 ng/μL), 10 μL of 10× CutSmart Buffer, 4 μL of Xho I enzyme, and 66 μL of nuclease-free water. Mix gently and then incubate at 37 °C for 30 min in a PCR thermal cycler to linearize the vector.
2. PCR amplification of the RUBY fragment: Set up two 50 μL PCR reactions in 0.2 mL PCR tubes using a high-fidelity DNA polymerase. For each reaction, add 25 μL of 2× PCR buffer for KOD FX, 4 μL of dNTPs, 2 μL of forward primer (5′-TATTATGGAGAAACTCGAGCTTATCTTTAATCATATTCCATAGT-3′), 0.8 μL of reverse primer (5′-ACAAATCTATCTCTCTCGAGATGGATCATGCGACCCTCGCCATGA-3′), 0.8 μL of template DNA, 0.5 μL of KOD FX, and 16.9 μL of nuclease-free water. Perform PCR according to the manufacturer’s recommended cycling conditions.
3. Purify the linearized vector and RUBY insert: Load the Xho I–digested pCAMBIA1300 backbone and PCR-amplified RUBY fragment onto a 1% (w/v) agarose gel prepared in 1× TAE buffer. Perform an agarose gel electrophoretic run at 120 V for 20 min. Visualize DNA bands using a gel imaging system, excise the target bands with a UV gel cutter, and transfer each gel slice into a 1.5 mL microcentrifuge tube. Purify DNA from gel slices using a commercial gel extraction kit according to the manufacturer’s instructions. Measure DNA concentration using a spectrophotometer and confirm fragment size by agarose gel electrophoresis.
4. Assemble the pCAMBIA1300-RUBY construct by seamless cloning: Assemble the linearized vector and RUBY insert using a ClonExpress MultiS One Step Cloning kit according to the manufacturer’s instructions. It contains 4 μL of 5× CE MultiS Buffer, 2 μL of Exnase MultiS, 2 μL of linearized pCAMBIA1300 backbone (200 ng), and 0.5 μL of RUBY insert (100 ng). Mix gently and incubate at 37 °C for 30 min, followed by cooling to 4 °C or immediate placement on ice.
5. Transform the assembled plasmid into Escherichia coli: Thaw 50 μL of chemically competent E. coli DH5α cells on ice. Add 5 μL of the assembly reaction directly to the competent cells and gently flick the tube to mix. Incubate the cells on ice for 30 min, heat-shock at 42 °C for 45 s, and return the cells to ice for 2 min. Add 700 μL of LB medium without antibiotics and incubate at 37 °C with shaking at 220 rpm for 1 h. Centrifuge the culture at 5000× g for 1 min, discard the supernatant, and resuspend the cell pellet in 200 μL of LB medium. Plate the suspension onto LB agar containing 50 μg/mL kanamycin and incubate the plates inverted at 37 °C overnight.
6. Verify positive clones: Pick single colonies and inoculate each into 5 mL of LB containing 50 μg/mL kanamycin in a 50 mL culture tube. Incubate cultures at 37 °C with shaking at 220 rpm overnight. Perform colony PCR using forward primer 5′-TCAACACATGAGCGAAACCCTAT-3′ and reverse primer 5′-ACAAATCTATCTCTCTCGAGATGGATCATGCGACCCTCGCCATGA-3′. Send PCR-positive clones for Sanger sequencing to confirm the correct insertion of the RUBY cassette. Store verified clones in 10% (v/v) glycerol at -80 °C.
Note: For soybean hairy root transformation, expression vectors carrying the RUBY reporter can be constructed following this protocol. Alternatively, a pre-existing RUBY vector with a functional expression cassette can be directly applied. In this protocol, the template used to amplify the RUBY fragment was derived from the original RUBY vector (35S:RUBY, Addgene plasmid #160908, https://www.addgene.org/160908/) [20].
B. Preparation of A. rhizogenes competent cells
1. Streak A. rhizogenes strain K599 from a -80 °C glycerol stock onto LB agar plates. Incubate the plates inverted at 28 °C in the dark for 48 h.
2. Pick a single colony and inoculate it into 5 mL of LB liquid medium. Culture the cells overnight at 28 °C with shaking at 200 rpm.
3. Transfer 2 mL of the overnight culture into 50 mL of fresh LB liquid medium in a 250 mL Erlenmeyer flask. Grow the culture at 28 °C with shaking at 200 rpm until the optical density at 600 nm (OD600) reaches 0.5–0.6.
4. Place the culture on ice for 30 min, then centrifuge at 5,000× g for 5 min at 4 °C. Discard the supernatant and gently resuspend the bacterial pellet in 10 mL of pre-chilled sterile 0.1 M NaCl solution.
5. Centrifuge the suspension again under the same conditions and resuspend the pellet in 1 mL of pre-chilled 20 mM CaCl2 solution. Resuspend 100 μL aliquots of competent cells in sterile 1.5 mL microcentrifuge tubes with glycerol to a final concentration of 10% (v/v). Immediately plunge the tubes into liquid nitrogen and then store them at -80 °C until use.
Note: Prepare A. rhizogenes K599 competent cells as described above or purchase commercially available competent cells for direct use.
C. Transformation of RUBY expression vector into A. rhizogenes
1. Thaw A. rhizogenes K599 competent cells on ice. Mix 50 μL of competent cells with 2 μL of pCAMBIA1300-RUBY plasmid DNA (approximately 500 ng) in a 1.5 mL microcentrifuge tube.
2. Incubate the tube on ice for 5 min, freeze the tube in liquid nitrogen for 5 min, then heat-shock it in a 37 °C water bath for 5 min. Immediately return the tube to ice for 5 min. Add 500 μL of antibiotic-free LB liquid medium to the tube. Incubate the cells at 28 °C with shaking at 200 rpm for 2–3 h to allow recovery.
3. Spread 200 μL of the recovered culture onto LB agar plates containing 50 μg/mL kanamycin and 50 μg/mL streptomycin. Invert the plates and incubate in the dark at 28 °C for 2–3 days.
4. Pick single colonies and verify the presence of the RUBY cassette or candidate gene by colony PCR. Inoculate PCR-positive colonies into 5 mL of LB liquid medium containing 50 μg/mL kanamycin and 50 μg/mL streptomycin. Culture at 28 °C with shaking at 220 rpm for 36 h.
5. Streak 100 μL of the overnight culture onto LB agar plates containing 50 μg/mL kanamycin, 50 μg/mL streptomycin, and 15 μg/mL acetosyringone. Incubate inverted at 28 °C in the dark for 24 h. Use the resulting bacterial cultures directly for soybean hairy root transformation.
6. (Optional) To preserve transformed A. rhizogenes, add sterile glycerol to a final concentration of 10% (v/v), mix gently, aliquot into sterile microcentrifuge tubes, and store at -80 °C.
Note: The pCAMBIA1300-RUBY vector described in this protocol can be used directly for soybean hairy root transformation as a visual transformation marker. Alternatively, based on experimental objectives, target genes or genome-editing cassettes can be inserted into the pCAMBIA1300-RUBY backbone for gene function analysis, provided that the RUBY reporter cassette remains intact and functional.
D. Soybean seed germination and seedling preparation
1. Select soybean cultivar Williams 82 and choose seeds with smooth surfaces and no visible lesions for hairy root transformation.
2. Surface-sterilize the seeds by chlorine gas. Prepare the sterilization setup by mixing 96 mL of NaClO with 4 mL of 37% HCl in a sealed desiccator. Place the seeds inside the desiccator and incubate for 12 h at room temperature.
Caution: Chlorine gas is highly toxic. The desiccator should be placed in a chemical fume hood.
3. After sterilization, sow the seeds evenly in nutrient soil in 17 cm × 17 cm soil trays. Place the trays in a controlled growth chamber set to a 14/10 h light/dark photoperiod at 25 °C with 65% relative humidity.
4. Grow seedlings for 4 days until they reach the stage shown in Figure 1B.
E. Establishment of transgenic soybean hairy root composite plants
1. Use a sterile surgical blade to create a wound at a 45° angle on the hypocotyl of 4-day-old soybean seedlings. Collect A. rhizogenes strain K599 carrying the pCAMBIA1300-RUBY vector and immediately apply the bacterial culture to the wound site (Figure 1C–G).
2. Place three sterile filter papers in a glass Petri dish and moisten them with 10 mL of sterile distilled water. Place the inoculated seedlings onto the filter papers with the wounded hypocotyls facing upward, cover the Petri dish to maintain humidity, and co-cultivate in the dark at 25 °C for 24 h (Figure 1H).
3. Transfer the seedlings to transparent plastic containers filled with moist vermiculite for hairy root induction. Place the containers in a tray and cover them with lids to maintain humidity. Grow the seedlings in a controlled growth chamber under a 14/10 h light/dark photoperiod at 25 °C with 65% relative humidity (Figure 1I).
4. Approximately 7 days post-inoculation, observe callus formation at the wound site. During this period, remove any adventitious roots that emerge to ensure clear hairy root development (Figure 1J, K).
Note: Non-transgenic adventitious roots can affect transformation efficiency. During the induction period, they are typically removed twice.
5. Approximately 10 days post-inoculation, observe distinct red hairy roots emerging from the wound site (Figure 1K). One to two days after the hairy roots appear, remove the moisture-retaining lid and continue cultivation under the same growth conditions.
Note: Ensure that the hairy roots have grown to at least 3–4 cm before removing the lid.
6. Continue to grow the hairy roots for an additional 5 days. Gently remove the roots from the vermiculite and rinse thoroughly with sterile distilled water to remove any adhering particles (Figure 1L, M). Use the clean transgenic hairy roots for downstream analyses, such as candidate gene expression assays, protein–protein interaction studies, metabolite profiling, or other functional experiments.
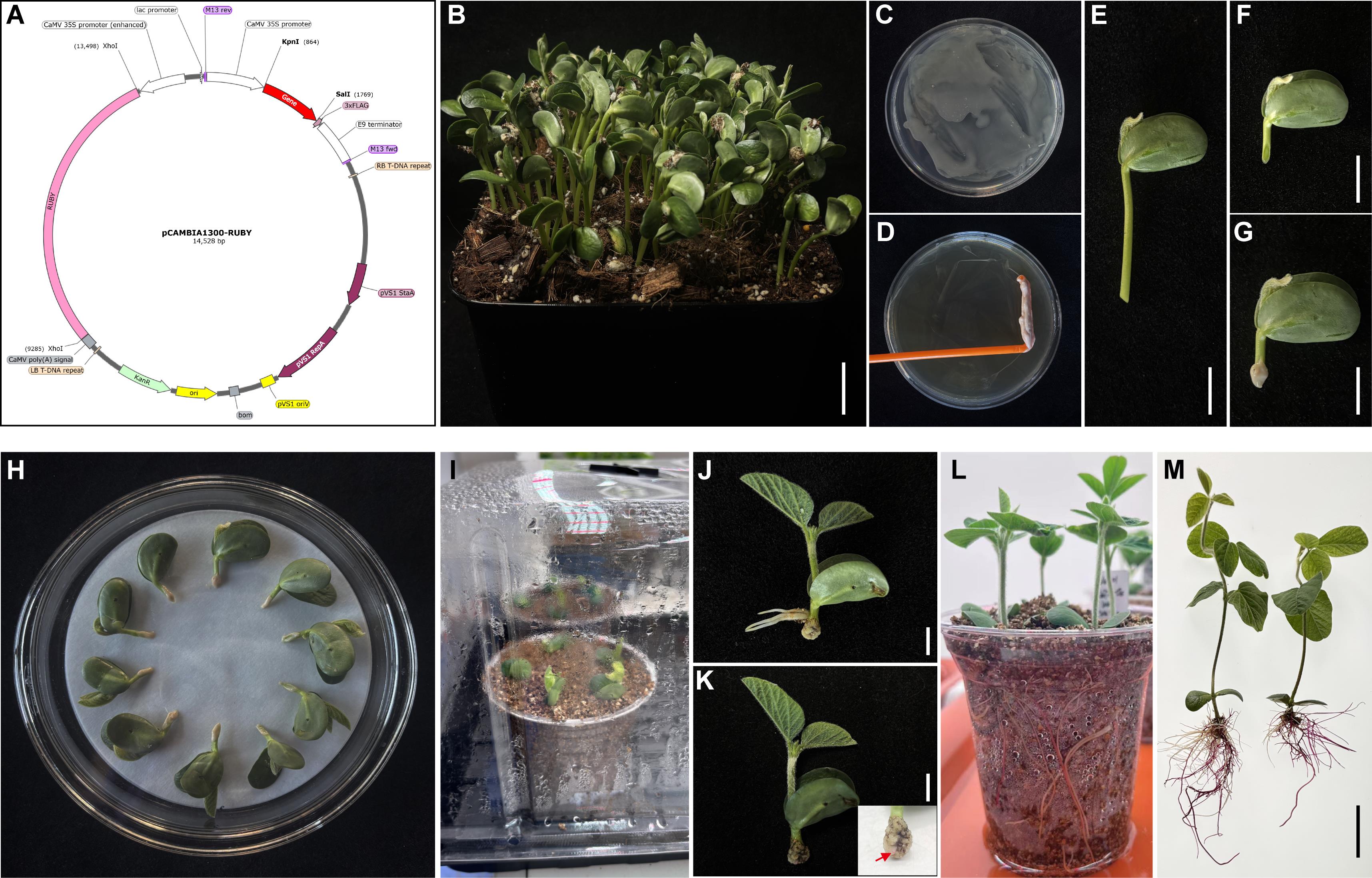

Figure 1. Generation of transgenic soybean hairy roots using the pCAMBIA1300-RUBY vector. (A) Schematic map of the pCAMBIA1300-RUBY binary vector, illustrating key elements: RUBY reporter gene for visual selection, driven by the cauliflower mosaic virus 35S (CaMV 35S) promoter and terminated by the CaMV polyA signal, Kpn I and Sal I restriction sites for cloning, kanamycin resistance gene (Kan^R) for bacterial selection, and T-DNA borders for plant integration. (B) Soybean seedlings grown in soil under controlled conditions (25 °C, 14/10 h light/dark) prior to transformation. Scale bar = 2 cm. (C) A. rhizogenes K599 carrying the pCAMBIA1300-RUBY vector was activated for 24 h. (D) The bacterial culture was concentrated using a sterile spreader. Scale bar = 1 cm. (E) Four-day-old soybean seedlings were excised at the base of the hypocotyl. Scale bar = 1 cm. (F) Hypocotyls were wounded at a 45° angle approximately 1 cm below the cotyledon node. Scale bar = 1 cm. (G) The wound sites were inoculated with A. rhizogenes K599. Scale bar = 1 cm. (H) Inoculated explants were placed on sterile filter paper in a glass dish with 10 mL of sterile water for co-cultivation in the dark for 24 h. (I) After co-cultivation, explants were transferred into vermiculite for hairy root induction and kept moist to prevent wilting. (J) During hairy root induction, adventitious (non-transformed) roots may form and should be carefully removed to select true hairy roots. Scale bar = 1 cm. (K) Callus formation at the infection site 7 days after induction. The enlarged inset shows the co-cultivation site with distinct white callus tissue; red arrows indicate RUBY-derived red pigmentation at approximately 10 days after induction and mark emerging positive hairy roots. Scale bar = 1 cm. (L) Fifteen days after induction, transgenic hairy roots fully developed, and RUBY expression was visible as red pigmentation. (M) Hairy roots are gently removed from vermiculite, washed with sterile water to remove debris, and are ready for downstream analyses. Scale bar = 2 cm.
Data analysis
Hairy root induction efficiency was calculated using the following formula:
where NRUBY-positive plants represents the number of plants producing at least one RUBY-positive hairy root, and Ntotal plants represents the total number of inoculated plants in each experiment.
The average number of positive roots per plant was calculated using only RUBY-positive plants and expressed as mean ± standard deviation (SD).
Each experimental repeat represents an independent biological replicate, with individual plants treated as independent transformation events. No data points were excluded from the analysis.
Using this analysis, induction efficiencies of approximately 90%–92.86% were consistently obtained across four independent repeats when assessed 20 days after induction, with an average of 3.85–5.80 RUBY-positive roots per plant (Table 1). These results demonstrate the robustness and reproducibility of the protocol.
All calculations and descriptive statistics were performed using Microsoft Excel. No specialized statistical software or computational expertise is required for data analysis in this protocol.
Table 1. Hairy root induction efficiency and RUBY-positive hairy root numbers in soybean
| Repeat | Total plants | RUBY-positive plants | Induction efficiency (%) | Average number of positive roots per plant |
|---|---|---|---|---|
| Rep 1 | 11 | 10 | 90.91 | 5.80 ± 2.35 |
| Rep 2 | 14 | 13 | 92.86 | 3.85 ± 1.82 |
| Rep 3 | 14 | 13 | 92.86 | 4.58 ± 1.56 |
| Rep 4 | 10 | 9 | 90.00 | 5.00 ± 2.45 |
Validation of protocol
To validate this protocol, four endogenous soybean genes—GmCHS7, GmC4H1, GmCHI4a, and GmChOMT—were cloned into the pCAMBIA1300-RUBY expression vector containing Kpn I and Sal I restriction sites within the multiple cloning site. The coding sequences of these genes were placed under the control of the Cauliflower Mosaic Virus 35S (CaMV 35S) promoter for constitutive expression in soybean hairy roots. Expression levels of the four genes were quantified by RT-qPCR using GmACT11 as the internal reference. Three biological replicates were included, and the empty pCAMBIA1300-RUBY vector (EV) served as a negative control. Relative gene expression was calculated using the 2^−ΔΔCT method. All four genes exhibited significantly higher expression levels compared with the EV control (Figure 2), demonstrating that this system supports effective transgene expression in soybean hairy roots and provides a robust platform for downstream functional or protein studies.
In addition, this protocol has been used and validated in the following research article:
• Wang et al. [22] Spatial Flavonoid Accumulation in Soybean Pericycle Restricts Phytophthora sojae Invasion. Preprint available at Research Square. (Supplementary Figure S7D–G).
As presented in Wang et al. [22], this protocol was applied to generate transgenic soybean hairy roots overexpressing disease-resistance genes. Using these roots, the invasion patterns of Phytophthora sojae were observed in relation to overexpressed resistance genes, and metabolomic analyses were performed to investigate the functional roles of these genes in soybean defense responses.
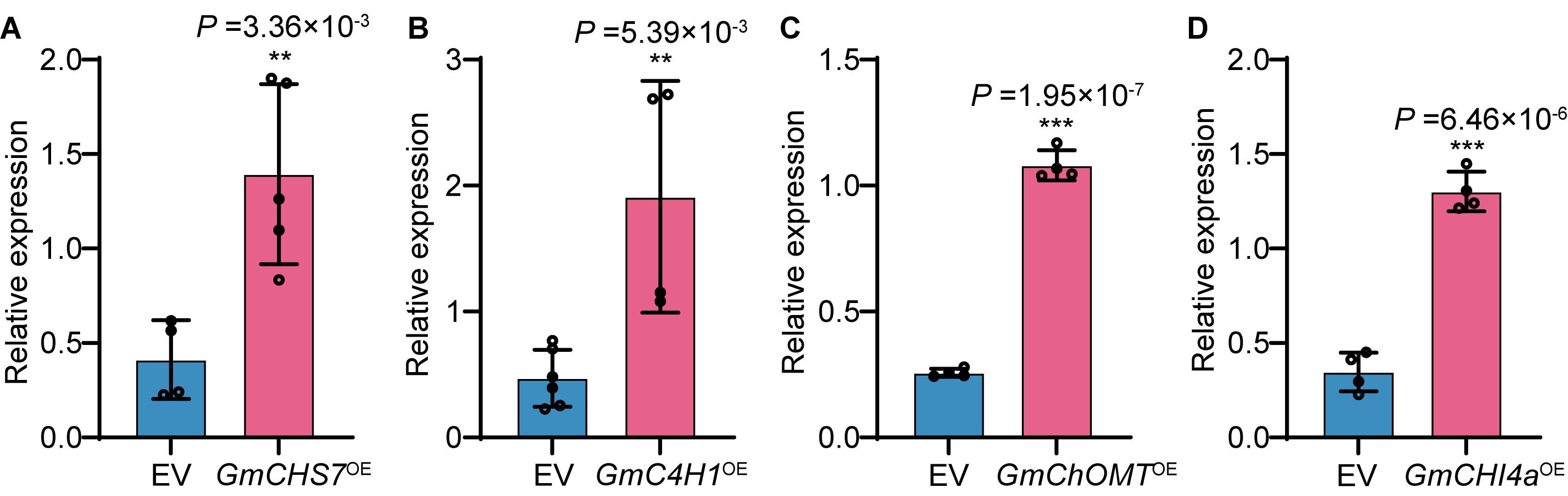
Figure 2. Relative expression levels of target genes in transgenic soybean hairy roots determined by RT-qPCR. Panels A–D show GmCHS7, GmC4H1, GmChOMT, and GmCHI4a, respectively. Data represent mean ± SD of three biological replicates, each consisting of two technical replicates. Statistical significance was determined by Student’s t-test. **p ≤ 0.01; ***p ≤ 0.001.
General notes and troubleshooting
General notes
1. Genotype dependency: Transformation efficiency may vary among soybean genotypes. While this method has achieved efficiencies exceeding 90% in cultivars such as Williams 82, HN66, and BX10 [19], and over 80% in recalcitrant wild soybean accessions (with at least 8 out of 10 plants producing positive roots in our trials), genotype-dependent effects cannot be entirely ruled out. Therefore, when applying this protocol to other genotypes, it is recommended to perform a preliminary test and adjust experimental conditions if necessary.
2. Selection of A. rhizogenes strain: A. rhizogenes strain K599 was selected for this protocol due to its documented high virulence and stability in soybean hairy root transformation. However, other strains may be suitable depending on the specific soybean genotype or experimental requirements. If alternative strains are employed, preliminary optimization of infection parameters is recommended to ensure maximal transformation efficiency.
3. Contamination control: Although this protocol does not require strictly aseptic conditions throughout the entire procedure, basic contamination control during the co-cultivation stage is recommended. Using sterile Petri dishes, sterile filter paper, and sterile water during co-cultivation can effectively reduce microbial contamination and improve the reliability of hairy root induction.
4. Flexibility of labware: Consumables such as plant growth trays, Petri dishes, or containers mentioned in this protocol can be substituted with functionally equivalent alternatives, provided they accommodate the seedling size and maintain appropriate humidity levels.
Troubleshooting
Problem 1: Overgrowth of Agrobacterium during co-cultivation.
Possible causes: Excessively high bacterial density or prolonged co-cultivation time.
Solutions: Reduce the volume or density of the bacterial suspension applied to the wound site, and limit co-cultivation to 24 h. If necessary, briefly rinse the seedlings with sterile water before transferring them to vermiculite to remove excess bacteria.
Problem 2: Low transformation efficiency.
Possible causes: Inefficient T-DNA delivery, reduced Agrobacterium virulence, or suboptimal physiological state of the explant.
Solutions: Use a virulent A. rhizogenes strain carrying the binary vector and inoculate with cultures in the exponential phase (OD600 = 0.5–0.6). Target the hypocotyl region ~1 cm below the cotyledonary node of healthy 4-day-old seedlings to maximize transformation efficiency. Remove adventitious roots during induction (typically twice) to improve the selection of positive roots. If efficiency remains low, alternative soybean genotypes may be tested.
Problem 3: Seedling death.
Possible causes: Excessive mechanical damage during wounding or dehydration during co-cultivation and hairy root induction.
Solutions: Minimize tissue damage during wounding. Maintain high humidity during co-cultivation by properly covering the plates. During hairy root induction, transfer seedlings to moist vermiculite and ensure the substrate is consistently moist but not waterlogged. Remove humidity covers only after hairy roots reach 3–4 cm in length.
Acknowledgments
J.H.Z. and Y.G. conceived and designed the protocol; Z.R.Z. and Q.S.W. performed the experiments and wrote the manuscript; J.H.Z. provided funding support. All authors read and approved the final manuscript. This work was supported by the Agriculture Science and Technology Major Project in China. This protocol is adapted from our previous work [22].
Competing interests
The authors declare no conflicts of interest.
References
- Byrne MC, Koplow J, David C, Tempé J, Chilton MD. (1983). Structure of T-DNA in roots transformed by Agrobacterium rhizogenes. J Mol Appl Genet. 2(2): 201–209. https://pubmed.ncbi.nlm.nih.gov/6875427/
- Bahramnejad, B., Naji, M., Bose, R. and Jha, S. (2019). A critical review on use of Agrobacterium rhizogenes and their associated binary vectors for plant transformation. Biotechnol Adv. 37(7): 107405. https://doi.org/10.1016/j.biotechadv.2019.06.004
- Gomes, C., Dupas, A., Pagano, A., Grima-Pettenati, J. and Paiva, J. A. P. (2019). Hairy Root Transformation: A Useful Tool to Explore Gene Function and Expression in Salix spp. Recalcitrant to Transformation. Front Plant Sci. 10: e01427. https://doi.org/10.3389/fpls.2019.01427
- Yang, Z., Gao, Z., Zhou, H., He, Y., Liu, Y., Lai, Y., Zheng, J., Li, X. and Liao, H. (2021). GmPTF1 modifies root architecture responses to phosphate starvation primarily through regulating GmEXPB2 expression in soybean. Plant J. 107(2): 525–543. https://doi.org/10.1111/tpj.15307
- Pistelli, L., Giovannini, A., Ruffoni, B., Bertoli, A. and Pistelli, L. (2010). Hairy Root Cultures for Secondary Metabolites Production. Adv Exp Med Biol. 698: 167–184. https://doi.org/10.1007/978-1-4419-7347-4_13
- Qin, Y., Wang, D., Fu, J., Zhang, Z., Qin, Y., Hu, G. and Zhao, J. (2021). Agrobacterium rhizogenes-mediated hairy root transformation as an efficient system for gene function analysis in Litchi chinensis. Plant Methods. 17(1): 103. https://doi.org/10.1186/s13007-021-00802-w
- Estrada-Navarrete, G., Alvarado-Affantranger, X., Olivares, J. E., Guillén, G., Díaz-Camino, C., Campos, F., Quinto, C., Gresshoff, P. M. and Sanchez, F. (2007). Fast, efficient and reproducible genetic transformation of Phaseolus spp. by Agrobacterium rhizogenes. Nat Protoc. 2(7): 1819–1824. https://doi.org/10.1038/nprot.2007.259
- Morey, K. J. and Peebles, C. A. M. (2022). Hairy roots: An untapped potential for production of plant products. Front Plant Sci. 13: e937095. https://doi.org/10.3389/fpls.2022.937095
- Kong, Q., Li, J., Wang, S., Feng, X. and Shou, H. (2023). Combination of Hairy Root and Whole-Plant Transformation Protocols to Achieve Efficient CRISPR/Cas9 Genome Editing in Soybean. Plants. 12(5): 1017. https://doi.org/10.3390/plants12051017
- Olhoft, P. M., Flagel, L. E., Donovan, C. M. and Somers, D. A. (2003). Efficient soybean transformation using hygromycin B selection in the cotyledonary-node method. Planta. 216(5): 723–735. https://doi.org/10.1007/s00425-002-0922-2
- Xu, H., Guo, Y., Qiu, L. and Ran, Y. (2022). Progress in Soybean Genetic Transformation Over the Last Decade. Front Plant Sci. 13: e900318. https://doi.org/10.3389/fpls.2022.900318
- Liu, S., Kandoth, P. K., Warren, S. D., Yeckel, G., Heinz, R., Alden, J., Yang, C., Jamai, A., El-Mellouki, T., Juvale, P. S., et al. (2012). A soybean cyst nematode resistance gene points to a new mechanism of plant resistance to pathogens. Nature. 492(7428): 256–260. https://doi.org/10.1038/nature11651
- Wang, F., Chen, H., Li, Q., Wei, W., Li, W., Zhang, W., Ma, B., Bi, Y., Lai, Y., Liu, X., et al. (2015). GmWRKY27 interacts with GmMYB174 to reduce expression of GmNAC29 for stress tolerance in soybean plants. Plant J. 83(2): 224–236. https://doi.org/10.1111/tpj.12879
- Ma, Z., Zhu, L., Song, T., Wang, Y., Zhang, Q., Xia, Y., Qiu, M., Lin, Y., Li, H., Kong, L., et al. (2017). A paralogous decoy protects Phytophthora sojae apoplastic effector PsXEG1 from a host inhibitor. Science. 355(6326): 710–714. https://doi.org/10.1126/science.aai7919
- Cheng, Y., Wang, X., Cao, L., Ji, J., Liu, T. and Duan, K. (2021). Highly efficient Agrobacterium rhizogenes-mediated hairy root transformation for gene functional and gene editing analysis in soybean. Plant Methods. 17(1): 73. https://doi.org/10.1186/s13007-021-00778-7
- Song, J., Tóth, K., Montes‐Luz, B. and Stacey, G. (2021). Soybean Hairy Root Transformation: A Rapid and Highly Efficient Method. Curr Protocol. 1(7): e195. https://doi.org/10.1002/cpz1.195
- Kereszt, A., Li, D., Indrasumunar, A., Nguyen, C. D., Nontachaiyapoom, S., Kinkema, M. and Gresshoff, P. M. (2007). Agrobacterium rhizogenes-mediated transformation of soybean to study root biology. Nat Protoc. 2(4): 948–952. https://doi.org/10.1038/nprot.2007.141
- Fan, Y. l., Zhang, X. h., Zhong, L. j., Wang, X. y., Jin, L. s. and Lyu, S. H. (2020). One-step generation of composite soybean plants with transgenic roots by Agrobacterium rhizogenes-mediated transformation. BMC Plant Biol. 20(1): 208. https://doi.org/10.1186/s12870-020-02421-4
- Ferguson, S., Abel, N. B., Reid, D., Madsen, L. H., Luu, T. B., Andersen, K. R., Stougaard, J. and Radutoiu, S. (2023). A simple and efficient protocol for generating transgenic hairy roots using Agrobacterium rhizogenes. PLoS One. 18(11): e0291680. https://doi.org/10.1371/journal.pone.0291680
- He, Y., Zhang, T., Sun, H., Zhan, H. and Zhao, Y. (2020). A reporter for noninvasively monitoring gene expression and plant transformation. Hortic Res. 7(1): 152. https://doi.org/10.1038/s41438-020-00390-1
- Sharifova, S., Prasad, K. V., Cheema, A. and Reddy, A. S. (2025). Genetically encoded betalain-based RUBY visual reporters: noninvasive monitoring of biological processes. Trends Plant Sci. 30(12): 1332–1345. https://doi.org/10.1016/j.tplants.2025.05.017
- Wang, Q. S., Dong, Z. M., Wang, H. Y., Zhang, Z. R., Ma, C. Y., Tan, X. W., Wang, Q. Q., Geng, Y. and Zhao, J. H. (2025). Spatial Flavonoid Accumulation in Soybean Pericycle Restricts Phytophthora sojae Invasion. bioRxiv. https://doi.org/10.64898/2025.11.30.691358
Article Information
Publication history
Received: Dec 30, 2025
Accepted: Feb 11, 2026
Available online: Feb 27, 2026
Published: Mar 20, 2026
Copyright
© 2026 The Author(s); This is an open access article under the CC BY-NC license (https://creativecommons.org/licenses/by-nc/4.0/).
How to cite
Zhang, Z., Wang, Q., Geng, Y. and Zhao, J. (2026). A Rapid and Visual Soybean Hairy Root Transformation Protocol Using the RUBY Reporter. Bio-protocol 16(6): e5635. DOI: 10.21769/BioProtoc.5635.
Category
Plant Science > Plant transformation > Agrobacterium
Molecular Biology
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








