- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
A Rapid and High-Recovery Extracellular Vesicle (EVs) Isolation Technique from Blood Samples
Published: Vol 16, Iss 5, Mar 5, 2026 DOI: 10.21769/BioProtoc.5632 Views: 24
Reviewed by: Alessandro DidonnaAnonymous reviewer(s)
Abstract
Extracellular vesicles (EVs) circulating in blood serve as non-invasive “liquid biopsies,” carrying molecular cargo that reflects the physiological and pathological state of distant cells. Their analysis is crucial for understanding disease mechanisms and discovering novel biomarkers. Clinically, blood EVs hold significant promise for early disease diagnosis, prognostic assessment, and monitoring treatment response in diverse areas such as organ transplantation, cancer, and neurological disorders. Current EV isolation techniques, beyond ultracentrifugation, include size exclusion chromatography (separation by size for high purity) and immunoaffinity capture (using antibodies for high specificity). Here, we present a simplified, rapid, and reproducible method for isolating EVs from small-volume blood samples. This protocol consistently yields a concentrated EV pellet covering 50–300 nm EVs, amenable to direct downstream analysis. Developed and validated in our laboratory using human, porcine, and murine blood samples, this method has proven instrumental in identifying EV-based biomarkers for predicting outcomes related to organ transplantation. The protocol’s adaptability and reliance on readily prepared, cost-effective reagents further enhance its utility. This scalable approach can be further integrated with subsequent purification or enrichment steps to optimize sample preparation for protein and nucleic acid assays.
Key features
• This method uses Ficoll-isolated plasma or centrifuged serum from any source, fresh to frozen.
• Protocol is easy to handle, fast, and has fewer technical details.
• 50–200 μL of plasma can be used to isolate enough EVs to run 5–8 western blot gels.
• Protocol acquires 50–300 nm EVs enriched with markers and nucleic acids.
• EVs isolated by this method can be further purified.
Keywords: EVs isolationGraphical overview
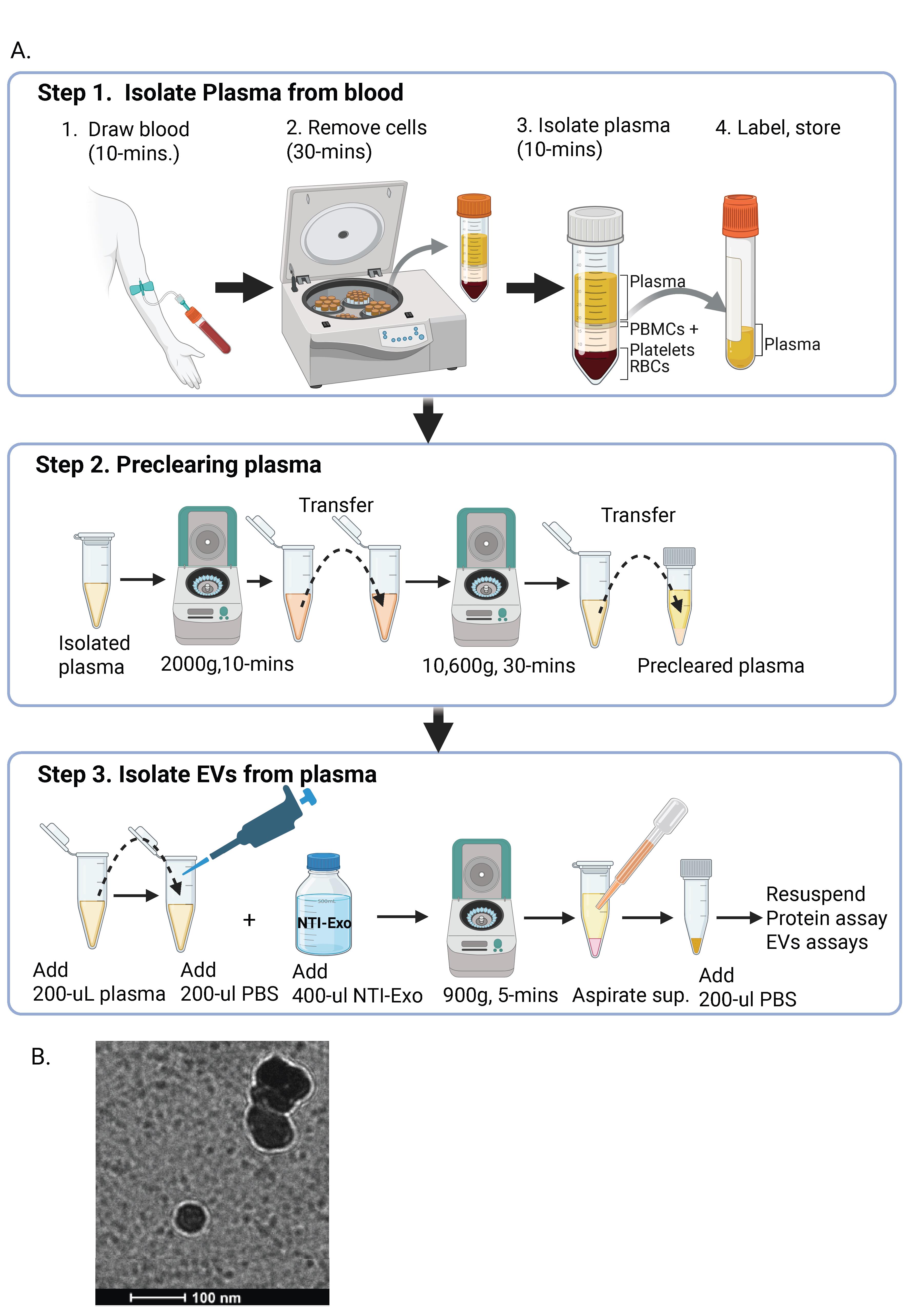
Graphical overview of extracellular vesicle (EV) isolation from plasma samples. (A) Flowchart of the isolation of extracellular vesicles from plasma, detailing plasma preparation and EV purification. (B) Transmission electron microscopy (TEM) image of EVs isolated from human plasma samples. Scale bar, 100 nm.
Background
Extracellular vesicles (EVs), once considered mere cellular debris, have emerged as critical mediators of intercellular communication, playing pivotal roles in both physiological processes and pathological conditions. These diverse nanoscale membrane-bound structures, including exosomes and microvesicles, are secreted by virtually all cell types and carry a rich cargo of proteins, lipids, and nucleic acids that reflect the state of their parent cells. This intrinsic biological function has positioned EVs as promising biomarkers for disease diagnosis, monitoring treatment efficacy, and predicting disease progression across a wide spectrum of medical conditions [1]. The rapid growth in EV research has greatly advanced our understanding of their biological role and potential clinical use. The ability to non-invasively sample and analyze patient-derived EVs offers an unparalleled opportunity for personalized medicine, moving beyond bulk tissue analysis to provide real-time insights into disease dynamics [2]. For instance, the presence of specific EV-encapsulated proteins or nucleic acids can serve as early indicators of disease onset, help to track the effectiveness of therapeutic interventions, or even predict patient response to particular treatments. This diagnostic and prognostic potential has fueled an intense drive to integrate EV analysis into routine clinical practice [3]. However, despite their immense promise, the widespread clinical application and comprehensive exploration of EVs face a significant bottleneck: the limitations of current isolation techniques. The inherent complexity and heterogeneity of biological fluids, coupled with the nanoscale dimensions of EVs, present formidable challenges. Several methods have traditionally been employed for EV isolation, each with its own set of advantages and disadvantages [4,5]. Ultracentrifugation (UC), long considered the gold standard, relies on differential centrifugation steps to progressively pellet EVs based on their size and density. While effective for isolating a broad range of EVs, UC is often criticized for its time-consuming nature, requirement for specialized equipment, potential for EV damage due to high G-forces, and the co-precipitation of contaminating proteins and aggregates, all of which necessitate skilled operators. Another common approach involves the use of size exclusion chromatography (SEC), which separates EVs based on their hydrodynamic radius as they pass through a porous matrix. SEC generally yields higher purity EV preparations and causes less mechanical stress compared to UC. However, SEC can suffer from lower recovery rates, particularly for smaller EVs, and can also be time-intensive, demanding careful column preparation and maintenance. Finally, commercial precipitation reagents offer a more convenient, often kit-based, approach to EV isolation. These methods typically involve the use of polymers that alter the solubility of EVs, causing them to aggregate and precipitate out of solution. While user-friendly and relatively rapid, precipitation methods can be prone to co-precipitation of non-EV components, leading to lower purity and potentially affecting downstream analyses. Furthermore, these reagents can sometimes interfere with the integrity or functionality of the isolated EVs.
Across all these established methods, recurring challenges persist for inconsistencies in yield and purity, compromises in EV integrity, and significant variations in reproducibility between laboratories. These issues are exacerbated when dealing with clinical samples of limited availability, such as those from pediatric patients, or when frequent, serial sampling is required for disease monitoring. The need for larger sample volumes, coupled with the labor-intensive nature of current techniques, often precludes their application in settings where only a few drops of blood are obtainable or where rapid turnaround times are crucial. The critical need for a more efficient, reliable, and accessible EV isolation method is underscored by the examples of disease-specific exosomal marker detection. Our laboratory’s investigations, including the identification of loss of LKB1 in transplant patients, SARS-CoV-2 spike protein in COVID-19 patients, and PD-L1 monitoring in immunotherapy recipients, all underscore the critical need for sensitive and accurate EV isolation from patient samples [6,7]. The lack of a robust, high-throughput, and low-volume compatible isolation technique has historically limited the potential for these exciting diagnostic and prognostic applications to translate from research laboratories into routine clinical practice.
This protocol is based on the understanding that an average EV content in human plasma is approximately 1010 per mL (ranging from 108 to 1013 per mL) [8,9]. This method, which has been validated and is routinely used, directly tackles persistent limitations by offering a rapid, user-friendly, high-recovery EV isolation technique for accessible clinical samples like blood [6]. Such an advancement is not merely incremental; it is fundamental to unlocking the full potential of EV-centric diagnostics, monitoring, and therapeutic strategies, ultimately propelling the frontiers of biomedical research and its application in personalized medicine.
Materials and reagents
Reagents
1. Commercial human serum (Sigma-Aldrich, catalog number: H4522)
2. Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: A886)
3. Dithiothreitol (DTT) (Sigma-Aldrich, catalog number: 3860-OP), store at -20 °C
4. Trizma base (Sigma-Aldrich, catalog number: RDD008)
5. EDTA (Sigma-Aldrich, catalog number: E6758)
6. Sodium chloride (NaCl) (Sigma-Aldrich, catalog number: S9888)
7. PEG (Sigma, catalog number: 81260)
8. Plasma [mice (C57BL/6J mice), or porcine (Yorkshire)], store at -20 °C
9. Anticoagulant: heparin (Sigma, catalog number: H3149)
10. 70% ethanol (for site disinfection, cotton-tipped applicators, and surgical gloves)
11. PierceTM BCA Protein Assay Kit (Thermo Scientific, catalog number: 23227)
12. RIPA lysis buffer
13. Sodium chloride (Sigma, catalog number: S5886)
14 Trizma base (Sigma, catalog number: 93352)
15. Deionized water (Milli-Q water)
16. Phosphate-buffered saline (PBS) (Sigma-Aldrich, catalog number: P4417)
17. Sodium acetate (JD Baker, catalog number:81260)
Primary antibodies used to identify EV markers
1. CD9 (BioLegend, catalog number: 312102)
2. CD63 (ProteinTech catalog number: 25682-1)
3. CD80 (ProteinTech, catalog number: 66406-1)
4. CD81 (Cell Signaling, catalog number: 56039)
5. Flotulin-1 (BioOrbit, catalog number: orb18698)
6. proteasome 20 s (Santa Cruz, catalog number: sc-271187)
7. NFKB (ProteinTech, catalog number: 66535)
8. Alix (Abcam, catalog number: ab88388)
9. TSG101 (Abcam catalog number: ab83)
10. CD63 (Santa Cruz Biotechnology, catalog number: sc-365604)
11. PD-L1 (Abcam, catalog number: ab213480)
12. CD73 (Cell Signaling Technology, catalog number: D7F9A)
13. ApoA1 (Cell Signaling, catalog number: 3350S)
14. Albumin (Cell Signaling, catalog number: 4929S)
15. Kappa light chain HRP (Thermo Fisher Scientific, catalog number: A18853)
Solutions
1. NTI-EXO solution (see Recipes)
Recipes
1. NTI-EXO solution
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Polyethylene glycol | 22.5% | 22.5 g |
| EDTA | 10 mM | 0.292 g |
| NaCl | 200 mM | 1.17 g |
| Tris-Cl | 200 mM | 2.42 g |
| Total | 100 mL |
Adjust to pH 7.2.
Laboratory supplies
1. 96-well flat-bottom plate (Thermo Scientific, catalog number: 21-377-832)
2. 1.5 mL Eppendorf tube (Fisher Scientific, catalog number: 50-274-4907)
3. Blood collection tubes (Fisher Scientific, catalog number: 22-040-134)
4. Cryovials (Simport Scientific, catalog number: T311-2)
5. 20 mL Syringe (BD, catalog number: 1625)
6. 0.2 µm sterile syringe filter (Thermo Scientific, catalog number: 723-2520)
7. Collection gear: 1 or 3 mL syringe with a 23–25 gauge needle
8. Polycarbonate ultracentrifuge tubes, 3 mL (Beckman Coulter, catalog number: 355635)
9. Surgical instruments: sharp dissecting scissors and thumb forceps
10. Graduated cylinder
11. Volumetric flasks
12. Sterile storage bottles
Equipment
1. Ultracentrifuge (Beckman Coulter, model: Optima XE-90) with a 70.1 TI fixed-angle titanium rotor
2. Nanosight NS300 (Software, Nanosight, NTA 3.3)
3. ExoView R200 platform (Software, ExoViewer 2.5.0)
4. pH meter (Thermo Scientific, model: Accumet AE150)
Procedure
Collection and processing of blood samples for EV isolation: This protocol details the collection and initial processing of blood samples from human patients, the Yorkshire porcine model used for transplant studies, and murine models, to obtain serum or plasma suitable for subsequent EV isolation. All procedures must adhere to approved institutional guidelines and ethical regulations.
General considerations:
1. Sterility: All steps involving sample collection and processing should be performed under sterile conditions to prevent contamination, especially if samples are destined for long-term storage or cell culture–based assays.
2. Temperature control: Maintain samples at appropriate temperatures (on ice, 4 °C, or -20/-80 °C for storage) as specified for each step to minimize degradation of analytes and EVs.
3. Safety: Always wear appropriate personal protective equipment (PPE), including a laboratory coat, gloves, and eye protection, when handling blood samples. Dispose of biohazardous waste according to institutional regulations.
A. Human sample collection and processing
Note: Ensure all human sample collection is performed under protocols approved by an accredited Institutional Review Board (IRB), such as the Institutional Review Board of St. Joseph’s Hospital and Medical Center committee. Obtain informed consent from all human participants prior to blood collection. Select appropriate blood collection tubes based on the desired final product (serum or plasma).
1. Collect venous blood samples from human patients into heparin tubes.
2. Immediately following collection, gently invert the tubes 5–10 times to ensure proper mixing with the anticoagulant. Centrifuge the whole blood at 2,000× g for 10 min at 4 °C (or room temperature, adjust as necessary based on downstream application). Carefully aspirate the supernatant plasma, taking care not to disturb the buffy coat or red blood cell layer.
3. Aliquot the separated plasma into cryovials in appropriate volumes to avoid repeated freeze-thaw cycles. Store aliquots at -20 °C or preferably -80 °C for long-term storage until EV extraction.
4. Procure pre-processed human serum from reputable suppliers. Store according to the manufacturer’s instructions, typically at -20 °C or -80 °C, upon receipt.
B. Porcine sample collection and processing
Note: All animal procedures must be conducted in strict accordance with protocols approved by the Institutional Animal Care and Use Committee (IACUC) of the facility. In our work, samples were collected from an ongoing study involving an organ care system experiment. Specific time points for blood collection should be strictly adhered to as per the main study protocol.
1. Collect porcine blood samples directly from the recycled reservoir of the organ care system to represent systemic circulation within the experimental setup.
2. Serum separation:
a. Allow collected blood to clot at room temperature for 30–60 min (or on ice for faster clotting if desired by specific research needs).
b. Centrifuge the clotted blood at 2,000× g for 10 min at 4 °C.
c. Carefully aspirate the supernatant serum, avoiding contact with the clot.
3. Aliquot the separated serum into cryovials and store them at -20 °C or preferably -80 °C until EV extraction.
C. Murine blood sample collection and processing
Note: In our work, all experimental protocols, including anesthesia, blood collection, and euthanasia, were approved by the Institutional Review Board of St. Joseph’s Hospital and Medical Center committee (or equivalent IACUC). All methods must be reported and performed in accordance with ARRIVE guidelines.
1. Anesthesia administration:
a. Anesthetize animals with a Ketamine/Xylazine cocktail (87 mg/kg).
b. Once initial anesthesia is achieved, clip the fur from the abdomen and restrain the animal.
c. Passively administer 1.0% isoflurane at 1 L/min via an inhalation system to maintain a sufficient level of anesthesia throughout the procedure.
d. Routinely verify anesthetic depth.
2. Surgical preparation for blood collection:
a. Perform a 3 cm abdominal midline incision to expose the inferior vena cava (IVC).
b. Maintain anesthesia and physiological parameters throughout this step.
3. Exsanguination (terminal blood collection):
a. Carefully visualize the IVC.
b. Perform transcardial puncture or direct IVC puncture/severance to allow for exsanguination.
c. Collect approximately 0.8 mL of blood into appropriate tubes (e.g., plain tubes for serum or anticoagulant-treated tubes for plasma).
4. Serum/plasma separation:
a. If collecting serum, allow blood to clot at room temperature for 30–60 min.
b. Centrifuge the whole blood (or clotted blood) at 2,000× g for 10 min at 4 °C to separate serum or plasma.
c. Carefully aspirate the supernatant serum/plasma.
5. Aliquoting and storage:
a. Aliquot the separated serum/plasma into cryovials.
b. Store aliquots at -20 °C or preferably -80 °C until EV extraction.
6. Following exsanguination, immediately euthanize mice by cervical dislocation while still under deep anesthesia, as approved by the IACUC protocol.
D. Post-collection handling (applicable to all sample types)
Note: If EV extraction is to be performed immediately, proceed directly to the EV isolation protocol (section E2) after serum/plasma separation. Prior to long-term storage, consider performing basic quality control checks on serum/plasma (visual inspection for hemolysis, protein concentration if required for normalization).
E. EV isolation using NTI-EXO precipitation
Note: This protocol describes a two-step method for the isolation of EVs from precleared serum or plasma samples using a polyethylene glycol (PEG)-based precipitation reagent (NTI-EXO). This method is designed to be rapid, reproducible, and cost-effective, particularly for small sample volumes (see the Materials section for the composition).
E1. Preparation of NTI-EXO reagent
1. Accurately weigh 22.5 g of PEG 6000 and transfer to a suitable beaker or flask.
2. Add approximately 60 mL of Milli-Q deionized water to the PEG and begin stirring on a magnetic stir plate to dissolve.
3. Concurrently, prepare the following stock solutions: 1.0 M Tris-HCl buffer, pH 7.2, and 0.5 M EDTA solution, pH 8.0 (adjust pH with NaOH).
4. To the dissolving PEG solution, add the following components while stirring: NaCl to a final concentration of 0.2 M; EDTA to a final concentration of 10 mM; and Tris-HCl (from a 2.0 M stock) to a final concentration of 200 mM (pH approximately 7.2).
5. Once all components are dissolved, adjust the solution volume to 100 mL with DI water.
6. Verify the pH of the final solution. It should be approximately 7.2 from the Tris buffering. Adjust with NaOH or HCl if necessary, though deviations are typically minimal if components were measured accurately.
7. Filter the entire NTI-EXO solution through a 0.2-µm sterile syringe filter or bottle-top filter into a sterile storage bottle. This step is critical to remove any potential contaminants and particulate matter that could interfere with downstream EV isolation or analysis.
8. Store the filtered NTI-EXO solution at 4 °C for up to one year. Label clearly with preparation date and initials.
E2. Preclearing of serum or plasma samples
Notes:
1. This step removes larger cellular debris, micro-vesicles, and larger protein aggregates that could co-precipitate with smaller EVs or interfere with downstream analysis.
2. Serum or plasma samples were collected and separated according to established protocols.
3. Samples should ideally be free of red blood cells.
1. Ensure that 250-μL serum or plasma samples are completely thawed if frozen. If samples contain any visible clots or significant particulate matter, an initial low-speed centrifugation (200× g for 5 min) can be performed, but this is usually covered by prior processing steps.
2. Transfer the cell-free serum or plasma into new, clean 1.5 mL microcentrifuge tubes.
3. Centrifuge the samples at 10,000 RCF for 30 min at 4 °C. This step is crucial for removing larger vesicles and apoptotic bodies.
4. Following centrifugation, carefully transfer the supernatant (precleared serum or plasma) to new, sterile 1.5 mL microcentrifuge tubes. Avoid disturbing the pellet at the bottom of the tube. This precleared supernatant is now ready for EV precipitation.
Note: The preclearing step is vital for the purity of the isolated EVs. Do not omit this step.
E3. EV isolation
Note: Polyethylene glycol (PEG) interacts with water molecules, effectively creating a volume exclusion effect that induces the precipitation of large macromolecules and EVs.
1. To each 1.5 mL microcentrifuge tube containing 200 μL of precleared serum or plasma, add 200 μL of sterile PBS. Mix gently by pipetting up and down.
2. Immediately add 400 μL of the prepared NTI-EXO reagent to each tube. The total reaction volume will be 800 μL.
3. Seal the tubes securely.
4. Place the tubes on a tube rotator and incubate at 4 °C for 30 min. Ensure continuous gentle rotation to facilitate EV precipitation.
5. Centrifuge the tubes at 750 RCF for 5–10 min at 4 °C. A visible pellet containing precipitated EVs should form at the bottom of the tube.
6. Gently aspirate and discard the supernatant without disturbing the EV pellet. This is a critical step; improper aspiration can lead to significant EV loss. One technique is to tilt the tube and aspirate away from the pellet.
7. Add 500 μL of sterile PBS to gently wash the EV pellet. Do not resuspend the pellet at this stage. Instead, gently flick the tube to ensure the PBS covers the pellet, or gently invert a few times. The wash step helps remove co-precipitated proteins and reagents.
8. Centrifuge again at 750 RCF for 5 min at 4 °C.
9. Carefully aspirate and discard the wash supernatant.
10. Add 200 μL of sterile PBS to the washed EV pellet. Vortex gently but thoroughly for 10–15 s to resuspend the EVs.
11. If the pellet does not fully solubilize after vortexing (a visible clump remains), cap the tube and place it at 4 °C overnight (12–16 h). This allows the pellet to rehydrate and dissolve slowly.
12. After overnight incubation, vortex again (10–15 s) and centrifuge any remaining large aggregates (2,000 RCF for 5 min) to obtain a clear EV suspension. The supernatant contains the isolated EVs.
E4. Alternative procedure (low-pH NTI-EXO acetate method)
1. In a 1.5 mL microcentrifuge tube, combine 100 μL of precleared serum.
2. Add 5 μL of 1 M sodium acetate, pH 4.5.
3. Add 60 μL of 5 M NaCl.
4. Add 35 μL of PBS and 200 μL of NTI-EXO reagent.
5. Mix gently and proceed with rotation and resuspension as described in steps E3.4–12.
F. Quantification of isolated EVs
Note: Quantification of total protein concentration of the isolated EV preparations is performed using a bicinchoninic acid (BCA) Protein Assay kit. This provides a common metric for comparative analysis between different EV samples.
1. Prepare working reagent according to the BCA kit manufacturer’s instructions.
2. Prepare a series of BSA standards of known concentrations to generate a standard curve.
3. Load 3 μL of each standard (0, 1, 2, 3, 4, 5, and 6 μg/well), duplicate sample, and blank (PBS) into the microplate wells.
4. Add 100 μL of working reagent to each well.
5. Incubate the plate at 37 °C for 25–30 min (or room temperature for 2 h, as per the kit’s instructions).
6. Measure the absorbance at 562 nm using the microplate reader.
7. Calculate protein concentration by comparing the absorbance to the standard curve.
Note: Protein concentration is an indirect measure of EV quantity. For more precise quantification, methods like nanoparticle tracking analysis (NTA) can be employed to determine particle concentration and size distribution.
G. Western blotting of isolated EVs
1. To prepare the samples for immunoblotting, dilute isolated EVs in 1× Laemmli buffer to a final concentration of 2 µg/µL, followed by denaturing at 95 °C for 10 min.
2. Resolve 20 µg of EVs on a NuPAGE 4%–12% Bis-Tris gel and transfer to a polyvinylidene difluoride (PVDF) membrane.
3. Rinse membranes with Milli-Q water and block for 30 min in 5% non-fat dry milk. Perform all washing and antibody dilution steps in 1× Tris-buffered saline with 0.1% Tween-20 (TBST).
4. Incubate membranes with primary antibodies against specific EV markers (CD9, CD63, CD80, CD81, Flotillin-1, Alix, and TSG101) and potential contaminants or functional targets (Proteasome 20s, NFκB, PD-L1, CD73, ApoA1, and Albumin)
5. Incubate membranes with appropriate HRP-conjugated secondary antibodies or kappa light chain HRP.
6. Visualize protein bands using SuperSignal West Pico PLUS Chemiluminescent Substrate and acquire digital images using an Odyssey-Fc Imager (LI-COR). Figure 1 shows a list of EV markers.
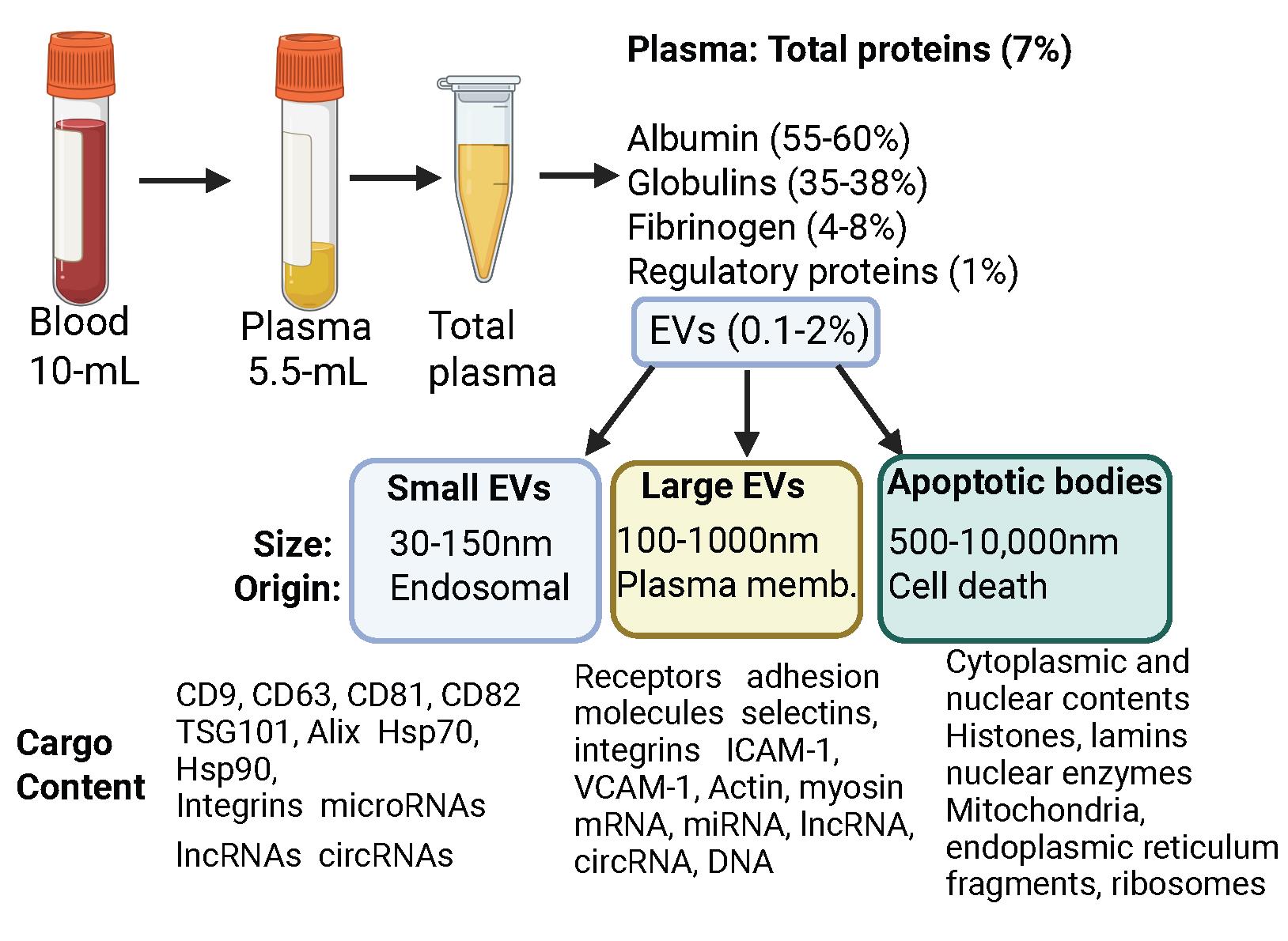
Figure 1. Overview of blood collection followed by plasma/serum collection. Following blood collection, plasma or serum is obtained, which contains a rich and varied pro tein content. Within this complex biofluid, EVs typically constitute a small fraction (0.1%–2% of total protein or volume, depending on patient condition). This figure depicts the diverse EV population, including small and large EVs, as well as distinct apoptotic bodies, highlighting their varied sizes and cellular origins.
H. Transmission electron microscopy (TEM)
1. Resuspend isolated exosome pellets in PBS to preserve morphology for imaging.
2. At a TEM facility, fix isolated EVs in 1 mL of 2.5% glutaraldehyde for 1 h at 4 °C.
3. Wash the fixed pellet three times in 0.5 mL of PBS and resuspend in PBS. For negative staining, adsorb fixed samples onto glow-discharged 200-mesh carbon-coated copper grids.
4. After brief drying, stain the grids with 3% phosphotungstic acid for 1 min, rinse with ultrapure water, and allow them to air dry completely.
5. Acquire images using a Talos L120C transmission electron microscope at an accelerating voltage of 120 kV. A TEM image of isolated EVs from human plasma is given in the Graphical overview.
Validation of protocol
Validation of the NTI-EXO method was established and published by Guerrero-Alba et al. [6]. A comparison with current technologies is shown in Figure 3H of that publication. The protocol’s robustness is evidenced by its ability to detect diverse biomarkers across various pathologies:
1. LKB1 suppression in EVs associated with human lung transplant rejection (Figure 6B).
2. SARS-CoV-2 spike protein in EVs from COVID-19 patients (Figure 6D).
3. PD-L1 expression in EVs from kidney transplant patient plasma (Figure 7D).
General notes and troubleshooting
Troubleshooting
Problem 1: Low purity/high contamination: High protein background in western blots, low EV-to-protein ratio, misleading particle counts by NTA (due to non-EV particles), and interference in downstream assays.
Possible causes: Suboptimal PEG reagent preparation or molecular weight leading to excessive co-precipitation. Inadequate removal of larger contaminants (cells, large debris) before PEG precipitation. Using old or not fully processed serum samples.
Solutions: This EV precipitation method offers significant advantages for discovering biomarkers in small-volume patient plasma samples. We consistently achieve reproducible results when using fresh plasma. EVs isolated via this protocol perform well in downstream analyses like Nanosight and Exoview assays. However, using older plasma samples, those subjected to multiple freeze-thaw cycles, or samples stored for extended periods at 4 °C, leads to degradation, high contamination, and low EV yield. Therefore, we recommend using relatively freshly acquired or appropriately stored plasma for optimal results. For omics discovery (proteomics, lipidomics, metabolomics), which demands highly pure EVs, this method provides superior recovery. It can be further integrated with size-exclusion chromatography (SEC) or density gradient ultracentrifugation for even greater EV purity
Problem 2: Aggregation of EVs: Broad or multiple peaks in NTA size distribution, visually clumpy or turbid samples, and reduced EV functionality due to masked surface proteins.
Possible causes: High local concentration of EVs and other macromolecules during precipitation. Presence of residual PEG acting as a crowding agent. Improper resuspension techniques, e.g., vigorous pipetting and vortexing. Long-term storage of concentrated EVs.
Solutions: Use gentle pipetting (slowly up and down), triturating, or gentle shaking instead of vigorous vortexing. Allow sufficient time for the pellet to fully dissolve. Use a neutral, iso-osmotic buffer, e.g., PBS, for resuspension. After adding resuspension buffer, allow the pellet to sit on ice for 30 min or overnight, with occasional gentle flicking or pipetting, to ensure full re-dispersion before centrifugation or further processing. Reduce PEG residue: Thorough washing steps (as described in Problem 1) are crucial to remove residual PEG.
Problem 3: Variable recovery and yield: Inconsistent EV particle counts or protein content across replicates or batches, difficulty obtaining sufficient material for downstream assays.
Possible causes: Suboptimal PEG concentration/molecular weight (too low for efficient precipitation). Insufficient incubation time for precipitation. Loss during washing or centrifugation steps (pellet not visible, too strong decanting). Batch-to-batch variation in serum/plasma composition.
Solutions: Carefully prepare NTI-EXO, use fresh plasma, and follow established protocols for consistent incubation time to ensure robust precipitation. Ensure the pellet is visible before decanting the supernatant. Slowly pour off the supernatant, avoiding disturbance of the pellet. If unsure, leave a small amount of supernatant (~50–100 μL), which can be removed during the wash stage/resuspension. Maintain strict consistency in all steps: temperature, mixing, centrifugation speeds and times, and resuspension volumes. For large volumes, consider using centrifugal filters (Amicon Ultra) to concentrate the sample before or after PEG precipitation, especially if using a subsequent SEC step.
Problem 4: Impact on EV integrity and functionality: Reduced EV uptake by target cells, altered EV surface marker expression, compromised RNA integrity, and negative results in functional assays.
Possible causes: Osmotic stress from high PEG concentration during precipitation. Direct interaction of PEG with EV membrane proteins/lipids. Physical stress during vigorous resuspension or centrifugation. Contaminating proteins interfering with EV function.
Solutions: Use gentle techniques to avoid mechanical stress. Use appropriate g-forces and ensure smooth acceleration/ deceleration ramps. Remove as much residual PEG as possible. If possible, use the minimum effective PEG concentration. For some sensitive applications, it might be better to slightly dilute concentrated EV samples rather than having them overly concentrated, reducing the potential for self-aggregation. SEC or density gradient UC: These methods are generally gentler on EVs and can remove harmful contaminants and PEG, preserving EV integrity and function more effectively than PEG alone.
Problem 5: Interference with downstream characterization and assays: Inaccurate NTA/DLS readings, smeared western blots, high background in ELISAs, poor RNA/DNA yield/quality from EVs, and compromised cell assays.
Possible causes: Co-precipitated proteins interfering with protein assays and obscuring EV-specific bands. Residual PEG interfering with enzyme activity, spectrophotometry, or cellular interactions. Aggregates skewing particle size and count measurements. Contaminating lipids or salts affecting nucleic acid extraction.
Solutions: Implement strategies from Problem 1 (pre-treatment, optimized PEG, thorough washing, secondary purification via UC or SEC) to eliminate contaminants. Ensure all wash steps are optimized to remove residual PEG. Dialysis can also be considered to remove PEG, though this can lead to EV loss. If residual PEG or protein contamination is unavoidable, strategic dilution of the sample for certain assays might reduce interference, though this also reduces EV concentration. For protein assays, consider kits designed for high-salt or detergent-containing samples if needed. Routinely test your isolated EVs for purity using a combination of methods (NTA, western blot for positive/negative markers, and total protein assay). Compare results with known contaminants.
Acknowledgments
We thank Norton Thoracic Institute and St. Joseph’s Hospital Medical Center for supporting this research. We thank Alexa Guerrero-Alba, Christin Poulson, Jesse Canez, and Kristina Sanborn for assisting with sample collection related to EVLP. This protocol was used in [6].
A.S., conceptualization, data acquisition, formal analysis, writing, review & editing. N.S., conceptualization, funding acquisition, supervision, data acquisition, formal analysis, validation, visualization, writing, review & editing. Both authors reviewed the manuscript.
Competing interests
We declare no competing interests.
References
- Zhang, X., Takeuchi, T., Takeda, A., Mochizuki, H. and Nagai, Y. (2022). Comparison of serum and plasma as a source of blood extracellular vesicles: Increased levels of platelet-derived particles in serum extracellular vesicle fractions alter content profiles from plasma extracellular vesicle fractions. PLoS One. 17(6): e0270634. https://doi.org/10.1371/journal.pone.0270634
- Bettio, V., Mazzucco, E., Antona, A., Cracas, S., Varalda, M., Venetucci, J., Bruno, S., Chiabotto, G., Venegoni, C., Vasile, A., et al. (2023). Extracellular vesicles from human plasma for biomarkers discovery: Impact of anticoagulants and isolation techniques. PLoS One. 18(5): e0285440. https://doi.org/10.1371/journal.pone.0285440
- Nieuwland, R. and Siljander, P. R. (2024). A beginner's guide to study extracellular vesicles in human blood plasma and serum. J Extracell Vesicles. 13(1): e12400. https://doi.org/10.1002/jev2.12400
- Figueroa‐Hall, L. K., Burrows, K., Alarbi, A. M., Hannafon, B. N., Hladik, C., Tan, C., Ramesh, R., Stewart, J. L., Risbrough, V. B., Paulus, M. P., et al. (2025). Comparison of Methods for Isolation and Characterization of Total and Astrocyte‐Enriched Extracellular Vesicles From Human Serum and Plasma. J Extracell Biol. 4(2): e70035. https://doi.org/10.1002/jex2.70035
- Holcar, M., Ferdin, J., Sitar, S., Tušek-Žnidarič, M., Dolžan, V., Plemenitaš, A., Žagar, E. and Lenassi, M. (2020). Enrichment of plasma extracellular vesicles for reliable quantification of their size and concentration for biomarker discovery. Sci Rep. 10(1): 21346. https://doi.org/10.1038/s41598-020-78422-y
- Guerrero-Alba, A., Bansal, S., Sankpal, A. N., Mitra, G., Rahman, M., Ravichandran, R., Poulson, C., Fleming, T. P., Smith, M. A., Bremner, R. M., et al. (2024). Enhanced enrichment of extracellular vesicles for laboratory and clinical research from drop-sized blood samples. Front Mol Biosci. 11: e1365783. https://doi.org/10.3389/fmolb.2024.1365783
- Rahman, M., Ravichandran, R., Sankpal, N. V., Bansal, S., Sureshbabu, A., Fleming, T., Perincheri, S., Bharat, A., Smith, M. A., Bremner, R. M., et al. (2023). Downregulation of a tumor suppressor gene LKB1 in lung transplantation as a biomarker for chronic murine lung allograft rejection. Cell Immunol. 386: 104690. https://doi.org/10.1016/j.cellimm.2023.104690
- Auber, M. and Svenningsen, P. (2022). An estimate of extracellular vesicle secretion rates of human blood cells. J Extracell Biol. 1(6): e46. https://doi.org/10.1002/jex2.46
- Johnsen, K. B., Gudbergsson, J. M., Andresen, T. L. and Simonsen, J. B. (2019). What is the blood concentration of extracellular vesicles? Implications for the use of extracellular vesicles as blood-borne biomarkers of cancer. Biochimica et Biophysica Acta (BBA) - Reviews on Cancer. 1871(1): 109–116. https://doi.org/10.1016/j.bbcan.2018.11.006
Article Information
Publication history
Received: Dec 29, 2025
Accepted: Feb 8, 2026
Available online: Feb 24, 2026
Published: Mar 5, 2026
Copyright
© 2026 The Author(s); This is an open access article under the CC BY-NC license (https://creativecommons.org/licenses/by-nc/4.0/).
How to cite
Sankpal, A. N. and Sankpal, N. V. (2026). A Rapid and High-Recovery Extracellular Vesicle (EVs) Isolation Technique from Blood Samples. Bio-protocol 16(5): e5632. DOI: 10.21769/BioProtoc.5632.
Category
Medicine
Molecular Biology
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link