现期刊物2026
卷册: 16, 期号: 11
生物信息学与计算生物学
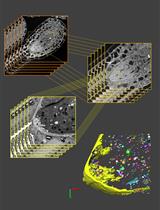
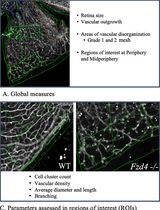
Computational Quantification of Mouse Retinal Vasculature Using ImageJ
基于 ImageJ 的小鼠视网膜血管图像定量分析
生物物理学
Versatile Dual Mounting Enables Larval Zebrafish Imaging Across Microscope Configurations
双向固定方法支持斑马鱼幼体在不同显微镜配置下成像
细胞生物学
In-Culture Antibody Capture Using Transient CHO Expression Systems
利用瞬时 CHO 表达系统在培养中捕获抗体
Comprehensive Protocol for Handling Human Small Airway Epithelial Cells (HSAECs) to Establish Air–Liquid Interface (ALI) Cultures With TEER-Based Barrier Integrity Assessment
人小气道上皮细胞气液界面培养及基于 TEER 的屏障完整性评估操作方案
发育生物学
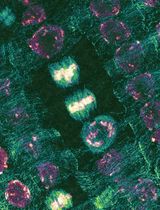
Quantitative Analysis of Cell and Tissue Shape During Mouse Cranial Neural Tube Closure
小鼠颅部神经管闭合过程中细胞与组织形态的定量分析
免疫学
A Reliable Method for Thawing Primary AML and CMML Mononuclear Cells to Preserve Viability and Function
保持原代 AML 和 CMML 单个核细胞活性与功能的可靠复苏方法
微生物学
Customizable High-Throughput Chemical Phenotyping of Root Bacteria
可定制的根际细菌高通量化学表型分析
分子生物学
Oligo(dT) Fluorescence In Situ Hybridization to Visualize the Poly(A) mRNAs in the Internal Tissues of Drosophila
利用 Oligo(dT) 荧光原位杂交观察果蝇内部组织中的 poly(A) mRNA
神经科学
Measuring Electrophysiological Activity in Acute Brain Slices, Spheroids, and Organoids Using 3D High-Density Multielectrode Arrays
利用三维高密度多电极阵列检测急性脑片、脑球体和脑类器官的电生理活动
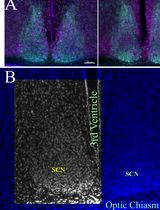
Histomorphometrical Analyses of the Mouse Suprachiasmatic Nucleus
小鼠视交叉上核的组织形态计量学分析
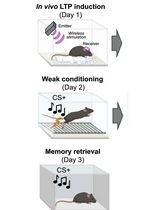
Optogenetic LTP Manipulation and Mathematical Modeling to Investigate Value Plasticity of the Instructive Signal in Mice
利用光遗传学 LTP 操控和数学建模研究小鼠指导性信号的价值可塑性
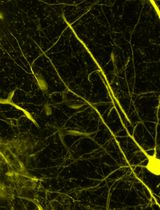
Detection of Target Molecules Within One-Millimeter-Thick Mouse Brain Slices by Using Peroxidase-Fused Nanobodies and Fluorochromized Tyramide-Glucose Oxidase Reaction
利用过氧化物酶融合纳米抗体和荧光化酪酰胺-葡萄糖氧化酶反应检测 1 mm 厚小鼠脑切片中的靶分子
植物科学
ROOT-ExM: Super-Resolution Imaging of Proteins in Arabidopsis Roots by Expansion Microscopy
ROOT-ExM:利用膨胀显微技术实现拟南芥根部蛋白质的超分辨率成像
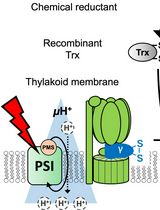
Evaluating Thioredoxin-Mediated CFoCF1 Reduction Using an In Vitro Thylakoid Assay
利用体外类囊体实验评估硫氧还蛋白介导的 CFoCF1 还原