- EN - English
- CN - 中文
A Novel Sequencing Method for Quantification of ZIKV RNA in Individual Cells
一种用于单细胞水平定量寨卡病毒RNA的新型测序方法
(*contributed equally to this work) 发布: 2026年03月20日第16卷第6期 DOI: 10.21769/BioProtoc.5645 浏览次数: 46
评审: Alessandro DidonnaLinda KangAnonymous reviewer(s)

相关实验方案
诱导型HIV-1库削减检测(HIVRRA):用于评估外周血单个核细胞中HIV-1潜伏库清除策略毒性与效力的快速敏感方法
Jade Jansen [...] Neeltje A. Kootstra
2025年07月20日 2494 阅读
Abstract
Single-cell RNA sequencing (scRNA-seq) is a powerful technique for exploring cellular heterogeneity and host–pathogen interactions. This protocol details the Zika virus (ZIKV)-targeted scRNA-seq workflow for preparing high-quality single-cell suspensions from the whole brain tissues of neonatal mice, high-quality single-cell sorting, cDNA reverse transcription, amplification, ZIKV enrichment and host transcriptome library preparation, and sequencing dataset integration in downstream analysis to complete the quantification of ZIKV RNA in individual cells.
Key features
• Preparation of high-quality single-cell suspensions from the whole brain tissues of neonatal mice.
• ZIKV-specific magnetic beads for using the ZIKV and host cell RNA capture.
• ZIKV enrichment and host transcriptome library construction, providing a framework for quantifying viral load within individual cells.
• Integration of viral enrichment and host transcriptomic datasets enables the visualization and quantification of ZIKV at single-cell resolution.
Keywords: Targeted single-cell RNA sequencing (靶向单细胞RNA测序)Graphical overview
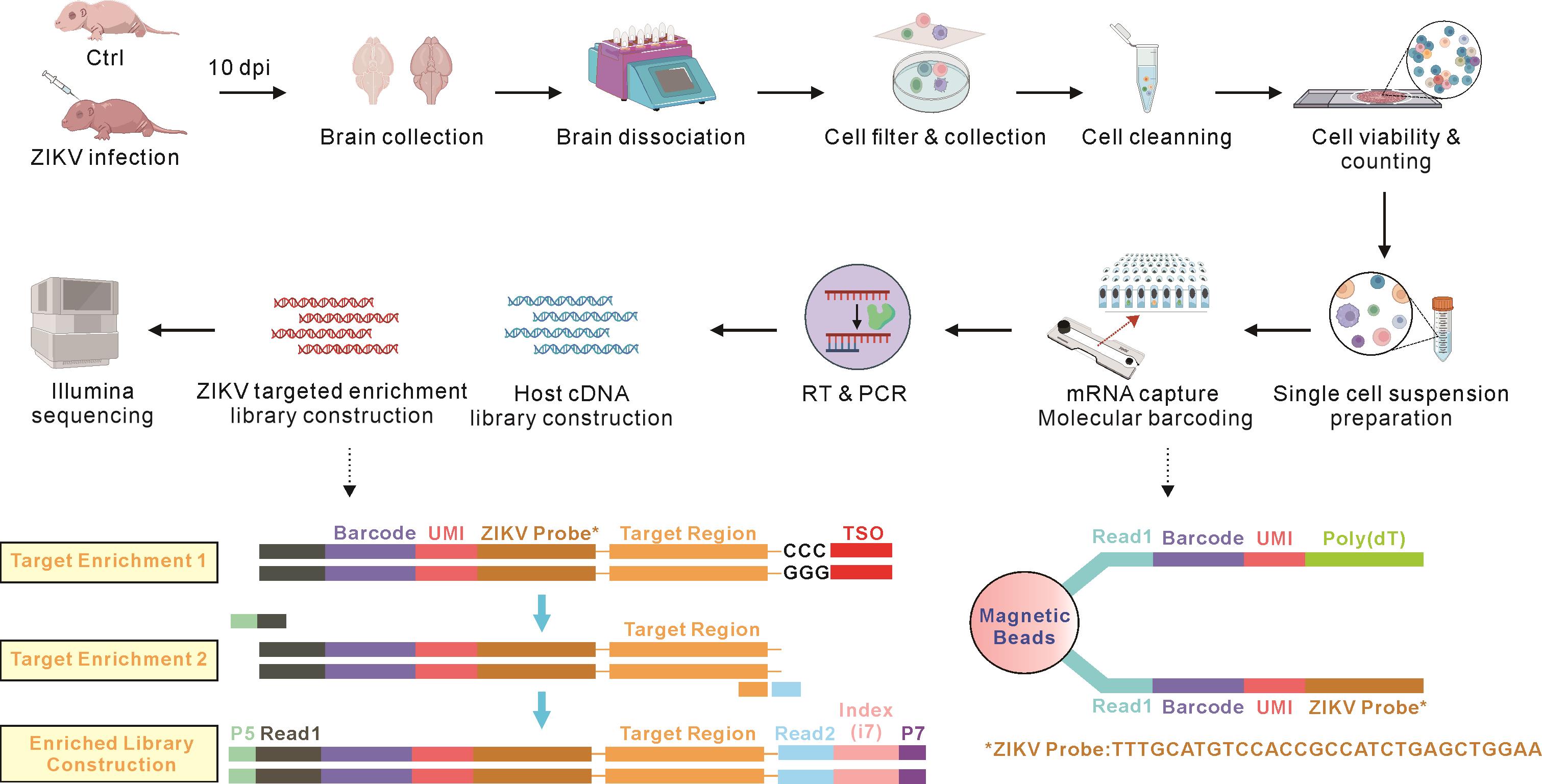
Working flow of Zika virus (ZIKV)-targeted single-cell RNA sequencing (scRNA-seq). Brain tissues of ZIKV-infected and uninfected (Ctrl) neonatal mice were collected after 10 days post-infection (dpi). Brain tissues were dissociated with Collagenase type-IV and filtered through a 200-mesh screen. Filtered cells were assessed for viability, density, and aggregation rate to prepare a qualified single-cell suspension. Eligible single cells and ZIKV-modified magnetic beads were loaded into a microfluidic chip to capture host and ZIKV mRNA, respectively. The mRNA underwent reverse transcription for library preparation. Following library quality control, separate host and ZIKV libraries were constructed, enriched, and sequenced on the Illumina platform. After data cleaning and filtering, the host and ZIKV datasets were integrated to analyze the cell characteristics and identify ZIKV-infected target cells.
Background
The Zika virus (ZIKV) infection seriously challenges global health by leading to neurological disorders, such as the Congenital Zika Syndrome (CZS) in newborns and the Guillain–Barré Syndrome (GBS) in adults. However, the mechanisms of ZIKV–host interaction remain unclear [1]. Single-cell RNA sequencing (scRNA-seq) is a critical tool for exploring cellular heterogeneity and pathogen–host interaction mechanisms at single-cell resolution [2,3]. However, the conventional scRNA-seq relies on poly(dT) primers to capture host RNA with poly(A) tails. ZIKV is an RNA virus that lacks a poly(A) tail, so it cannot be captured by poly(dT). Here, we provide a comprehensive protocol for ZIKV quantification in individual cells using the ZIKV-targeted scRNA-seq.
Materials and reagents
Biological materials
1. SMGC-1 ZIKV strains isolated from a traveler with a recent travel history to Fiji and Samoa, who was identified in Shenzhen, Guangdong Province, China (GenBank No. KX266255)
2. BALB/cAnNCrl neonatal mice (purchased from Beijing Vital River Laboratory Animal Technology Co., Ltd.)
Reagents
1. Collagenase type-IV (Sigma, catalog number: C4-BIOC)
2. DNase I (Roche, catalog number: 10104159001)
3. PBS (Thermo Fisher, catalog number: 14190094)
4. AO/PI staining solution (Counter Star, catalog number: 13200066)
5. SCellive® Cell Debris Removal kit (SCellive®, catalog number: 13200066)
6. Red blood cell (RBC) lysis buffer (Invitrogen, catalog number: 00-4333-57)
7. FocuSCOPE Single-Cell Multiomics mRNA × ZIKA Virus Detection kit (Singleron Biotechnologies, catalog number: 1002152-5)
Note: This kit was customized by modifying ZIKV-specific primers (5'-TTTGCATGTCCACCGCCATCTGAGCTGGAA-3') on the capturing beads.
8. Anhydrous ethanol (Merk, catalog number: 1.00983)
9. PBS-TWEEN® tablets (Merk, catalog number: 524653)
10. Cell lysis buffer (Merk, catalog number: 43-045)
11. QIAcardTM FTATM wash buffer (Merk, catalog number: WHAWB120204)
Solutions
1. Tissue digestion buffer (see Recipes)
2. Debris removal working solution (see Recipes)
Recipes
1. Tissue digestion buffer
| Reagent | Volume |
|---|---|
| PBS | 320 μL |
| Collagenase | 80 μL |
| DNase | 140 μL |
Store at 4 °C.
2. Debris removal working solution (for brain tissues) [4]
| Reagent | Volume |
|---|---|
| Debris removal buffer | 2.7 mL |
| Dilution buffer | 0.3 mL |
Laboratory supplies
1. Scissors (YuYan Instruments, catalog number: Y15103)
2. Finnpipette F3 (Thermo Scientific, catalog numbers: 4640040, 4640050, 4640060)
3. Finntip (Thermo Scientific, catalog numbers: 94052100, 94052200, 94052300, 94052410)
4. Countstar chamber slide (Count Star, catalog number: CO010101)
5. 60 mm × 15 mm tissue-culture treated culture dishes (Corning, catalog number: CLS430166)
6. Cell strainer, 200 screen mesh (Sangon Biotech, catalog number: F513442)
7. 15 mL centrifuge tubes (Corning, catalog number: CLS430791)
8. 1.5 mL microtubes (Corning, catalog number: AXYMCT150CS)
9. 1.5 mL low-binding microcentrifuge tubes (Biopioneer, catalog number: CNT-1.5FL)
Equipment
1. Digital heating shaking dry bath (Thermo Scientific, catalog number: 88880028)
2. Centrifuge 5920 R (Eppendorf, catalog number: 5948000093)
3. Countstar Rigel S5 (Count Star, catalog number: IN030301)
4. Inverted Microscope BX53 (Olympus, catalog number: BX53F2C)
5. Automated thermal cycler (Applied Biosystems, catalog number: A31491)
6. DynaMagTM-2 Magnet (Invitrogen, catalog number: 12321D)
Software and datasets
1. R software (free download from https://cran.r-project.org/bin/windows/base/) (version 4.3.1, 2023-10-31)
2. RStudio (free download from https://posit.co/download/rstudio-desktop/) (version 2023.03.1 + 446)
3. CeleScope (version 2.0.6, Singleron Biotechnologies) for barcode extraction, UMI counting, and read-level quality control
4. STAR (version 2.6.1a) for read alignment to the mouse reference genome (GRCm38) and the ZIKV reference genome (GenBank accession no. KU321639.1)
5. FeatureCounts (version 2.0.1, subread package) for gene-level quantification
6. Seurat (version v4.1.1) and DoubletFinder (version 2.0.3) R packages for normalization, integration, clustering, and doublet detection
7. MAST (version v1.18.0) R package for the analysis of differentially expressed genes (DEGs)
8. ggplot2 (version 3.4.0) for visualization
Procedure
文章信息
稿件历史记录
提交日期: Dec 9, 2025
接收日期: Feb 18, 2026
在线发布日期: Mar 5, 2026
出版日期: Mar 20, 2026
版权信息
© 2026 The Author(s); This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/).
如何引用
Hao, M., Wang, Y., Du, D., Yang, W., Guo, Q., Tang, M., Liu, L., Yang, W., Liu, Y., Luo, C., Chen, J., Wang, P., An, J. and Zhou, Y. (2026). A Novel Sequencing Method for Quantification of ZIKV RNA in Individual Cells. Bio-protocol 16(6): e5645. DOI: 10.21769/BioProtoc.5645.
分类
微生物学 > 微生物-宿主相互作用 > 病毒
细胞生物学 > 单细胞分析 > 流式细胞术
系统生物学 > 转录组学 > RNA测序
您对这篇实验方法有问题吗?
在此处发布您的问题,我们将邀请本文作者来回答。同时,我们会将您的问题发布到Bio-protocol Exchange,以便寻求社区成员的帮助。
Share
Bluesky
X
Copy link