- EN - English
- CN - 中文
Tandem RNA and Protein Extraction: A Platform for Maximizing the Use of Limited Ex Vivo Tissue Samples
RNA与蛋白质串联提取方法:最大化利用有限离体组织样本的实验平台
(*contributed equally to this work) 发布: 2026年03月20日第16卷第6期 DOI: 10.21769/BioProtoc.5643 浏览次数: 54
评审: Alberto RissoneKaustav MukherjeeAnonymous reviewer(s)

相关实验方案
髓系肿瘤中用于生殖系 DNA 分离的皮肤成纤维细胞标准化培养:一种从床旁到实验室的实用方法
Parampreet Kour [...] Pulkit Rastogi
2025年10月05日 1475 阅读
深海无脊椎动物组织保存的比较方案:DNA/RNA Shield 与液氮在高质量核酸双重提取中的应用对比
Ana S. Gomes [...] Olivier Laroche
2025年11月20日 1388 阅读
Abstract
Human tissue samples represent the gold standard for obtaining clinically relevant and translatable insight into disease processes that in vitro systems cannot fully reproduce. However, patient-derived samples are often limited in size and availability, limiting the number of downstream assays that can be performed. To maximize the use of invaluable human samples, we present a protocol for the tandem extraction of high-quality RNA and protein from the same tissue section. This method has been optimized for 15–30 mg tissue sections, enabling more experimental conditions and technical replicates, while minimizing intrasample variability associated with heterogeneous tissues. This protocol also avoids potentially hazardous solvents present in phenol-chloroform-based methods such as TRIzol, providing a safer and more accessible workflow without compromising biomolecule integrity. This protocol was developed and validated using atherosclerotic plaque tissue from carotid endarterectomy, a very challenging tissue type to work with due to extensive calcification, necrosis, and limited surgical availability. We have also validated this method using mouse aortic tissue and cultured THP-1 cells, demonstrating its versatility across sample input types. As this protocol relies on standard column-based RNA extraction kits and commonly available reagents for protein precipitation and extraction, this methodology is widely accessible and easy to implement as a standard, streamlined workflow.
Key features
• Advances ex vivo models of human disease (e.g., atherosclerosis) by maximizing the use of patient-derived tissue samples.
• Minimizes patient variability by generating directly comparable RNA and protein readouts.
• Minimizes the number of patient samples required and maximizes the output from each sample.
• Stabilizes RNA integrity from tissue without impacting the quality of proteins derived from the same sample.
Keywords: RNA extraction (RNA提取)Graphical overview
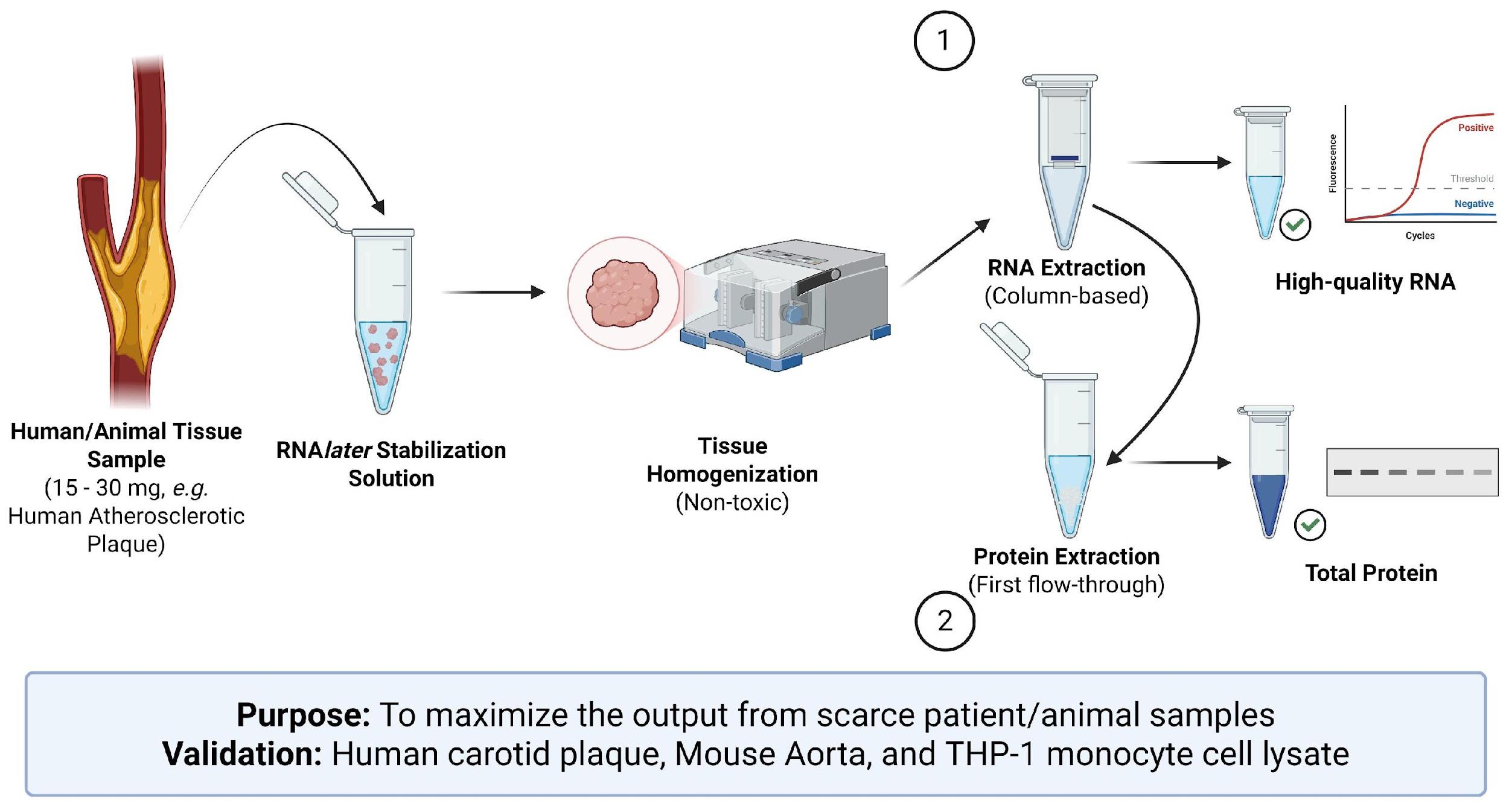
Background
Human tissue samples provide strong clinically relevant insight into disease processes that in vitro and in vivo systems cannot fully replicate. However, they pose some experimental challenges, such as heterogeneity and limited sample availability. Similarly, in vitro models, including cultured 3D models, possess their own limitations, as they do not recreate the integrated physiological environment of human disease. Animal models remain essential for mechanical and therapeutic studies, but translational issues often arise due to species-specific differences. For example, though human and murine macrophage subtypes share broad functional similarities, they diverge in important clinically relevant pathways, such as species-specific activation patterns of the PPARγ pathway [1]. Therefore, patient-derived tissue remains the most reliable source of directly translatable information.
The limited size and availability of patient samples restrict the number of downstream analyses that can be performed. This scarcity means that researchers often must be strategic about which assays to perform, given the limited number of samples available. To address this challenge, this protocol enables the tandem extraction of high-quality RNA and protein from the same tissue sample. This approach maximizes the usage of valuable human specimens while reducing intrasample variability, which may arise from sampling heterogeneous tissues.
This method is optimized for small tissue sections (15–30 mg), allowing more experimental conditions and technical replicates to be included. The use of RNAlater stabilizes RNA and protein, preserving integrity for one week at room temperature, one month at 4 °C, or indefinitely at -20 °C or lower [2]. Unlike common commercial methods like TRIzol, this workflow avoids toxic organic solvents like phenol and chloroform.
We developed and validated this protocol primarily with ex vivo–treated atherosclerotic plaque tissue from carotid endarterectomy (CEA). This tissue type provides substantial challenges because carotid plaques are often highly calcified due to long-term inflammation, necrosis, mechanical stress, aging, and osteogenic pathways, which are hallmarks of atherosclerotic disease. CEA tissue is also often limited in size and surgical availability. The use of small 15–30 mg (3 × 3 × 3 mm) sections for both RNA and protein could address some of these limitations. These small pieces of carotid tissue also remain metabolically active when cultured ex vivo for at least 24 h and are reactive to stimuli over this timeframe. This allows for the ex vivo treatment and screening of compounds of interest directly in human atherosclerotic tissue. The possibility of using small pieces of tissue and extracting both protein and RNA from them maximizes experimental outputs.
In addition to human CEA samples, we confirm the suitability of this protocol for use in mouse aortic tissue and THP-1 cells, and we expect this workflow to be adaptable to a wide range of tissue and sample types with minimal optimization (primarily in tissue volume and homogenization), because it relies on standard column-based RNA extraction kits and commonly available laboratory reagents. As a result, the protocol is widely accessible and compatible with most laboratory settings, providing an opportunity to maximize experimental readouts in most settings and a cheaper alternative to commercially available kits such as the Allprep (Qiagen) and Ambion (Thermo Fisher Scientific) kits. It also provides a layer of RNA stabilization, negating the need for liquid nitrogen snap freezing and tissue grinding for RNA extraction from tissue.
While several protocols have been described for precipitating total protein from various sources, including RNA extraction kit flowthrough [3–6], the protocol described here offers several distinct advantages. It avoids the use of hazardous organic compounds such as trichloroacetic acid, acetone, methanol, or chloroform. By avoiding these harsh organic compounds and avoiding washing with 96%–100% ethanol (as described in [6]), this protocol avoids stripping the proteins completely of hydration, facilitating easier downstream solubilization. Furthermore, by using only the first flowthrough of the RNA-extraction procedure for protein precipitation, samples can be processed in smaller volumes and tube sizes, reducing reagent and plasticware consumption.
Materials and reagents
Biological materials
1. Human tissue sample, e.g., carotid endarterectomy (approximately 3 × 3 × 3 mm in size or 15–30 mg in weight)
2. Animal tissue sample, e.g., snap-frozen mouse aorta (tissue samples of 15–30 mg, including small organs, can be processed whole if the tissue thickness does not exceed 5 mm)
3. Cell lysate, e.g., THP-1 monocytes
Reagents
1. β-mercaptoethanol, 100 mL (Sigma-Aldrich, catalog number: M3148)
2. Laemmli buffer and DTT, 8 mL (Cell Signaling Technology, catalog number: 7722S)
3. Ethanol 200-proof (Fisher Scientific, catalog number: CCE7023)
4. TRK lysis buffer (EZNA Extraction Kit) (Omega BIO-TEK, catalog number: R6834-02)
5. For tissue samples: Invitrogen RNAlater stabilization solution, 100 mL (ThermoFisher, catalog number: AM7020) (fresh or ex vivo–treated samples) or Invitrogen RNAlater-ICE tissue transition solution (ThermoFisher, catalog number: AM7030) (snap-frozen samples)
Solutions
1. 70% ethanol solution (see Recipes)
2. TRK lysis buffer with 2% β-mercaptoethanol (see Recipes)
Recipes
1. 70% ethanol solution
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Ethanol | 70% | 70 mL |
| Ultrapure water | - | 30 mL |
Store at 4 °C.
2. TRK lysis buffer with 2% β-mercaptoethanol
| Reagent | Final Concentration | Quantity or volume |
|---|---|---|
| TRK lysis buffer (or equivalent) | - | 9.8 mL |
| β-mercaptoethanol | 2% | 200 µL |
Store at 4 °C. Stable for one month.
Laboratory supplies
1. Column-based total RNA extraction kit (e.g., EZNA Total RNA Kit I, Omega BIO-TEK, catalog number: R6834-02)
2. 1.5 mL centrifuge tubes (Greiner, catalog number: 616201CIS)
3. 2 mL homogenizer tubes and caps (Sarstedt, catalog numbers: 72.609 and 65.716.004, respectively)
4. 50 and 15 mL Falcon tubes (Greiner, catalog numbers: 210270CIS and 188261CIS, respectively)
5. Pipette filter tips (P2–P1000)
6. Serological pipette strips (P5–P25)
7. 5 mm stainless steel beads (Qiagen, catalog number: 69989)
Equipment
1. Refrigerated centrifuge with a relative centrifugal force of at least 14,000× g
2. Bead homogenizer (QIAGEN, model: TissueLyser II)
3. Heat-block capable of 100 °C (Eppendorf, model: ThermoMixer_F2.0)
4. Fridge (4 °C)
5. Freezer (-20 °C and -80 °C)
6. Falcon tube and centrifuge tube racks
7. Nanodrop spectrophotometer (DeNovix DS-11 Spectrophotometer)
8. Pipettes (P2–P1000)
9. Stainless-steel tweezers (Merck, catalog number: 22435-U)
10. Scales (capable of measuring with milligram-level accuracy)
Procedure
文章信息
稿件历史记录
提交日期: Dec 19, 2025
接收日期: Feb 11, 2026
在线发布日期: Mar 4, 2026
出版日期: Mar 20, 2026
版权信息
© 2026 The Author(s); This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/).
如何引用
Kennedy, C., Millar, B., Conroy, L. J., Marai, M., Barry, M., O’Donohoe, M., Belton, O., Brennan, E., Godson, C. and de Gaetano, M. (2026). Tandem RNA and Protein Extraction: A Platform for Maximizing the Use of Limited Ex Vivo Tissue Samples. Bio-protocol 16(6): e5643. DOI: 10.21769/BioProtoc.5643.
分类
分子生物学 > RNA > RNA 提取
分子生物学 > 蛋白质 > 分离
细胞生物学 > 组织分析 > 组织分离
您对这篇实验方法有问题吗?
在此处发布您的问题,我们将邀请本文作者来回答。同时,我们会将您的问题发布到Bio-protocol Exchange,以便寻求社区成员的帮助。
Share
Bluesky
X
Copy link