- EN - English
- CN - 中文
A Simple and Cost-Effective Method for Generating Spheroids From Triple-Negative Breast Cancer Cell Line (MDA-MB-231)
一种简单且低成本的三阴性乳腺癌细胞系(MDA-MB-231)球状体构建方法
发布: 2026年03月20日第16卷第6期 DOI: 10.21769/BioProtoc.5641 浏览次数: 73
评审: Elena A. OstrakhovitchLivia E. SimaAnonymous reviewer(s)

相关实验方案
采用 Davidson 固定液和黑色素漂白法优化小鼠眼组织切片的免疫组化染色
Anne Nathalie Longakit [...] Catherine D. Van Raamsdonk
2025年11月20日 1624 阅读
Abstract
Breast cancer (BC) is the most frequently diagnosed malignancy in women and a leading cause of cancer-related mortality worldwide. Current clinical management relies on molecular classification—based on estrogen receptor (ER), progesterone receptor (PR), HER2, and Ki67 expression—to guide prognosis and therapy. Triple-negative breast cancer (TNBC), which lacks ER, PR, and HER2 expression, represents 15%–20% of cases and is characterized by aggressive behavior, early recurrence, and a paucity of targeted treatment options. These challenges underscore the urgent need for improved preclinical models that better recapitulate tumor biology to accelerate therapeutic discovery. While conventional monolayer (2D) cultures have contributed significantly to cancer research, they fail to mimic critical features of the three-dimensional (3D) tumor microenvironment (TME), thereby limiting clinical translation. To address this gap, 3D spheroid models have emerged as a powerful intermediary, more accurately replicating in vivo conditions such as cell–cell and cell–matrix interactions, nutrient and oxygen gradients, and the development of hypoxic cores. These features make spheroids a physiologically relevant platform for studying complex processes like metastasis, drug resistance, and treatment response. Here, we present a robust, simple, and cost-effective protocol for generating uniform 3D spheroids. Our method enables consistent monitoring of spheroid formation and growth over time, with quantitative, image-based size analysis to ensure reproducibility and scalability. Designed for flexibility, the protocol is broadly applicable across diverse cell types, effectively bridging the gap between traditional 2D cultures and complex in vivo studies. By providing an accessible and reliable model of the 3D TME, this protocol opens new avenues for high-throughput drug screening, mechanistic studies of tumor progression, and the advancement of personalized medicine strategies in breast cancer and beyond.
Key features
• Employs a cost-effective, lab-made agarose coating to create non-adherent surfaces in standard 96-well plates.
• Optimized for reproducible spheroid formation from the aggressive MDA-MB-231 triple-negative breast cancer cell line.
• Requires a brief orbital shaking step to promote homogeneous cell aggregation and formation of a single central spheroid per well.
• Generates measurable spheroids within 96 h using only basic cell culture equipment and an orbital shaker.
• Provides a clear workflow from spheroid formation to quantitative size analysis, using freely available (ImageJ) and widely used (GraphPad Prism) software.
Keywords: Triple-negative breast cancer (三阴性乳腺癌)Graphical overview
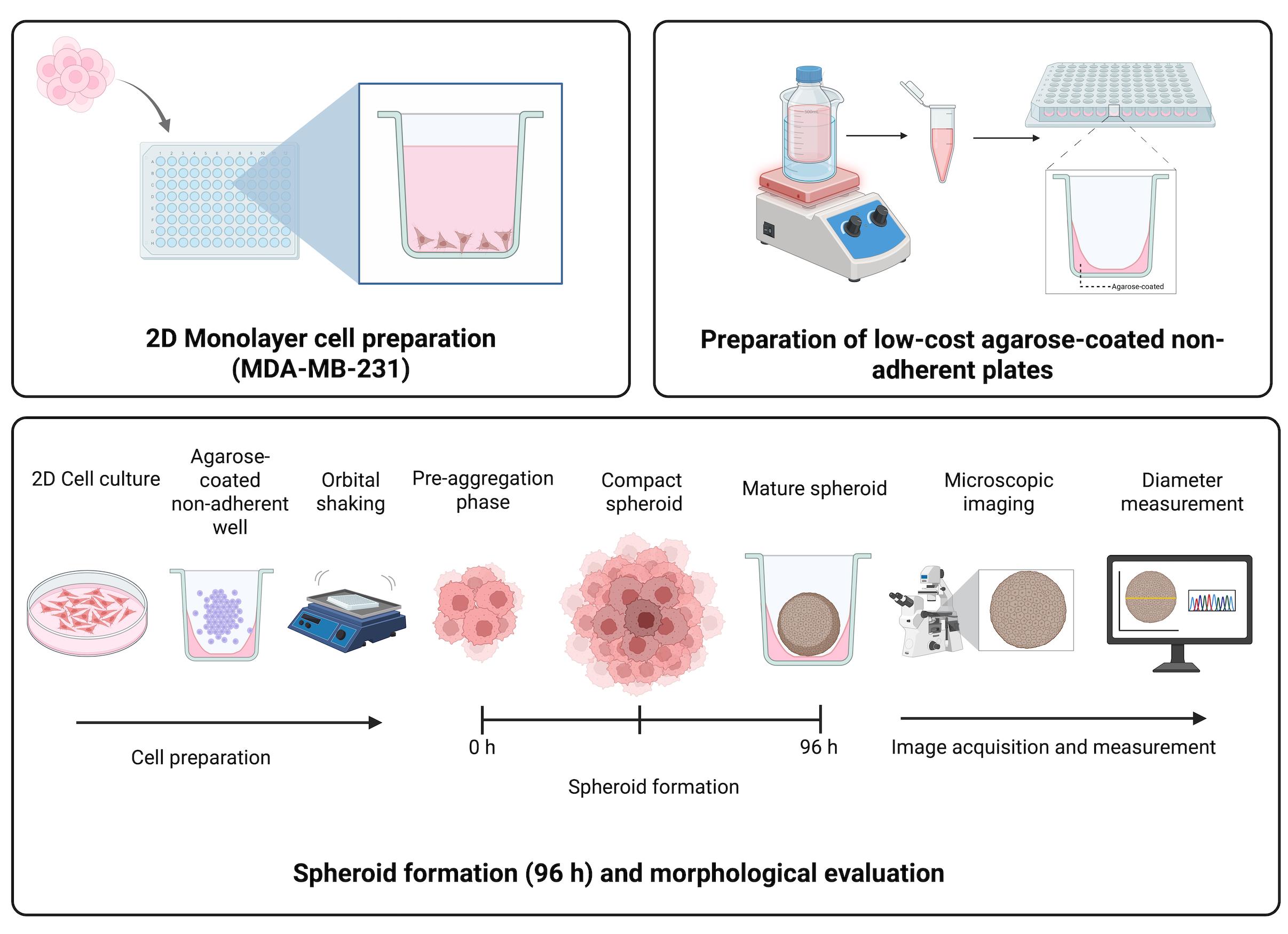
Background
Breast cancer (BC) remains the most diagnosed malignancy in women globally and the leading cause of cancer-related deaths [1]. BC is classified into molecular subtypes based on the expression of estrogen receptor (ER), progesterone receptor (PR), human epidermal growth factor receptor 2 (HER2), and the proliferation marker Ki67 [2,3]. These subtypes dictate prognosis and guide therapeutic strategies, with hormone receptor-positive (ER+/PR+) and HER2-enriched tumors benefiting from targeted therapies. Among these, triple-negative breast cancer (TNBC)—defined by the absence of ER, PR, and HER2 expression—represents 15%–20% of all breast cancers and is associated with aggressive clinical behavior, early recurrence, and limited treatment options [4]. Due to its lack of targetable receptors, TNBC patients primarily rely on chemotherapy, highlighting the urgent need for novel therapeutic approaches.
For decades, monolayer (2D) cell cultures have been the cornerstone of cancer research, enabling foundational discoveries in cell proliferation, migration, and drug screening [5]. However, these models fail to recapitulate the three-dimensional (3D) architecture and tumor microenvironment (TME) of in vivo tumors, leading to discrepancies in drug responses and mechanistic studies [6]. To bridge this gap, 3D spheroid models have emerged as a powerful tool, better mimicking the complexity of solid tumors [7]. These models incorporate cell–cell and cell–matrix interactions, polarization and spatial organization, oxygen and nutrient gradients, metabolic heterogeneity, and hypoxic cores that resemble in vivo conditions. As a result, spheroids provide a more physiologically relevant platform for studying tumor invasion and metastasis, drug resistance mechanisms, therapeutic screening, and personalized medicine approaches [8,9].
Recent advances in scaffold-based and scaffold-free spheroid generation techniques have improved reproducibility and scalability [10,11]. Among scaffold-free methods, the use of non-adherent surfaces is a cornerstone. Agarose coating is a well-established technique for creating non-adherent surfaces, promoting cell aggregation by minimizing substrate attachment. Previously described methodologies include coating standard multi-well plates with a thin layer of agarose [12, 13] and using agarose micromolds to create arrays of spheroids with uniform size and shape [14]. These approaches are valued for their simplicity, low cost, and compatibility with standard laboratory equipment. Other prevalent methods include the hanging drop technique, commercially available ultra-low attachment (ULA) plates with covalently modified hydrogel surfaces, and more complex microfluidic systems [10,11].
The spheroid generation method presented here, utilizing agarose-coated plates, aligns with this tradition of simple, accessible tools. Its primary advantages are universal applicability, minimal equipment requirements, and very low cost per sample, making it ideal for labs initiating 3D culture work or for high-volume exploratory studies. However, when compared to other published methods, the proposed protocol has inherent limitations. While ultra-low attachment plates offer superior reproducibility and higher throughput for screening assays, and microfluidic systems provide exquisite control over the microenvironment and dynamic perfusion, our agarose-coating method may exhibit more variability in spheroid size and shape between wells.
Despite these limitations, the protocol offers a simple, cost-effective, and universally applicable strategy to harness the power of 3D cell culture. Its ability to produce reliable in vitro models that accurately mimic key in vivo conditions remains a crucial advantage. We anticipate that the widespread adoption of this foundational protocol will enhance the predictive power of preclinical drug screening and serve as a robust tool for personalized therapy cancer research, particularly in settings where resource-intensive technologies are not available.
Materials and reagents
Biological materials
1. MDA-MB-231 cell line (ATCC, catalog number: CRM-HTB-26)
Reagents
1. Dulbecco’s modified Eagle medium (DMEM) (Merck, catalog number: D5648) or RMPI media
2. L-glutamine (Gibco, catalog number: 20530-081)
3. Amphotericin B (Sigma-Aldrich, catalog number: A2942)
4. Fetal bovine serum (FBS) (BioWest, catalog number: BIO-S1400)
5. Bovine calf serum (BCS) (BioWest, catalog number: S0400-500)
6. Penicillin-streptomycin (Gibco, catalog number: 15120122)
7. Trypsin (Sigma-Aldrich, catalog number: T4799-5G9)
8. Ethylenediaminetetraacetic acid (EDTA) (J.T. Baker, catalog number: 8993-01)
9. Sodium chloride (NaCl) (J.T. Baker, catalog number: 3624-01)
10. Sodium phosphate, 7-hydrate, crystal (J.T. Baker, catalog number: 3824-01)
11. Monobasic potassium phosphate crystal (J.T. Baker, catalog number: 3824-01)
12. Potassium chloride (KCl) (J.T. Baker, catalog number: 3040-01)
13. Non-adherent 96-well plates
14. Agarose (Invitrogen, catalog number: 16500-100)
15. MilliQ water sterile water (J.T. Baker, catalog number: 4220-20)
16. Sodium bicarbonate (NaHCO3) (J.T. Baker, catalog number: 3506-01)
17. Sodium phosphate (NaH2PO4) (J.T. Baker, catalog number: 3818-01)
18. Monopotassium phosphate (KH2PO4) (J.T. Baker, catalog number: 3246-01)
Laboratory supplies
1. Pipette tips: 1,000 μL (Uniparts, catalog number: 51131), 200 μL (Uniparts, catalog number: 51121Y), 10 μL (Axygen, catalog number: T-300)
2. MF-Millipore 0.22 μm MCE membrane, 47 mm (Merck, catalog number: GSWP04700)
3. Nitrile gloves (Kirkland Signature, catalog number: B0CJ4JMTQQ)
4. Culture tube, 15 mL (Uniparts, catalog number: 34117F)
5. Cell culture dish, 60 mm × 15 mm (Nest, catalog number: 705001)
6. Cell culture plate, 96 wells, flat bottom, tissue culture (TC) grade (Nest, catalog number: 701001)
7. Conical-bottom centrifuge tubes, 50 mL (Uniparts, catalog number: 32117F)
8. Conical-bottom centrifuge tubes, 15 mL (Uniparts, catalog number: 34117F)
9. Microcentrifuge tubes, 1.5 mL (Axygen, catalog number: MCT-150-C)
10. Serological pipettes, 2 mL (Corning, catalog number: 4486)
Solutions
1. Culture medium for MDA-MB-231 cells (DMEM complete medium) (see Recipes)
2. 1.5% agarose solution (see Recipes)
3. 1× PBS solution (see Recipes)
4. 0.04% Trypsin-EDTA (see Recipes)
Recipes
1. Culture medium for MDA-MB-231 cells (DMEM complete medium)
| Reagent | Final concentration | Quantity |
|---|---|---|
| DMEM powder | — | As required |
| Sodium bicarbonate (NaHCO3) | 3 g/L | 3 g |
| FBS | 5% (v/v) | 50 mL |
| BCS | 5% (v/v) | 50 mL |
| Penicillin–streptomycin | 1% (v/v) | 10 mL |
| Amphotericin B | 0.05% (v/v) | 500 μL |
| L-glutamine (200 mM) | 0.05% (v/v) | 500 μL |
| Sterile distilled water | — | To 1 L |
a. Dissolve DMEM powder in sterile distilled water and stir until completely dissolved.
b. Add 1.5 g of NaHCO3.
c. Adjust the pH to 7.4 by carefully adding drops of NaOH (1 M) or HCl (1 M), as needed.
d. Under a laminar flow hood, supplement the medium with the following components: FBS, BCS, 1% penicillin–streptomycin, 0.05% Amphotericin B, and 0.05% L-Glutamine 200 mM.
Note: This medium supports the growth of MDA-MB-231 cells under standard culture conditions. After preparation, the complete medium should be sterilized by filtration (0.22 μm MCE membrane in a vacuum filtration system) and stored at 4 °C for up to 4 weeks.
2. 1.5% agarose solution
| Reagent | Final concentration | Quantity |
|---|---|---|
| Agarose | 1.5% (w/v) | 1.5 g |
| Triple-distilled water | — | To 100 mL |
a. Add 1.5 g of agarose to 100 mL of triple-distilled water in a glass flask.
b. Sterilize the solution by autoclaving for 15 min.
Note: This agarose solution is used to generate non-adherent surfaces in standard 96-well plates. The prepared solution should be sterilized by autoclaving and used fresh or maintained at ~60–70 °C until dispensing.
3. 1× PBS solution
| Reagent | Final concentration | Quantity |
|---|---|---|
| NaCl | 137 mM | 8.0 g |
| KCl | 2.7 mM | 0.2 g |
| Disodium phosphate (Na2HPO4) | 10 mM | 1.44 g |
| Monopotassium phosphate (KH2PO4) | 1.8 mM | 0.24 g |
| Triple-distilled water | — | To 1 L |
a. Autoclave a glass flask containing 1 L of triple-distilled water and allow it to cool down at room temperature.
b. Add the following salts to the flask: NaCl, KCl, Na2HPO4, and KH2PO4.
c. Dissolve the salts using a magnetic stirrer.
d. Calibrate the pH meter and adjust the solution to pH 7.4 by slowly adding 1 M NaOH or 1 M HCl, as needed.
e. Remove the stir bar, filter the solution through a 0.22 μm membrane under sterile conditions, and store at 4 °C until use.
Note: Alternatively, a commercial PBS solution may be used.
4. 0.04% Trypsin-EDTA
| Reagent | Final concentration | Quantity |
|---|---|---|
| Trypsin | 0.04% (w/v) | 390 mg |
| EDTA | 0.037% (w/v) | 370 mg |
| NaCl | — | 85 mg |
| PBS 1× | — | To 1 L |
a. Start with 1 L of PBS 1×.
b. Dissolve 390 mg of trypsin and 85 mg of NaCl in 10 mL of PBS 1×.
c. Add 370 mg of EDTA to the remaining PBS 1× and mix until dissolved.
d. Combine with the trypsin–NaCl solution and stir until fully mixed.
e. Filter the solution through a 0.22 μm membrane under sterile conditions.
f. Aliquot and label the solution properly before storing at -20 °C until use.
Note: This trypsin-EDTA solution is used for cell detachment. The prepared solution should be sterilized by filtration (0.22 μm) and stored at -20 °C until use.
Equipment
1. Centrifuge (PowerSpinTM, model: C856)
2. Micropipettes (Axygen, AP-10, AP-100, and AP-1000): 0.5–10 μL single-channel pipettor (Axygen® Axypet®, catalog number: AP-10), 10–100 μL single-channel pipettor (Axygen® Axypet®, catalog number: AP-100), and 100–1,000 μL single-channel pipettor (Axygen® Axypet®, catalog number: AP-1000)
3. pH meter (Apera, model: PH700)
4. Orbital shaker (Benchmark, model: BT302)
5. Digital stirring hot plate (Thermo Scientific, model: SP131015Q)
6. Inverted microscope (Carl Zeiss, model: 37081)
7. Laminar flow cabinet (Thermo Fisher Scientific, model: 1340)
8. Incubator for cell culture (Thermo Fisher Scientific, model: 3422)
9. Neubauer chamber (Mariendfeld, catalog number: 0610010)
10. Microwave oven (standard laboratory or domestic units, 600–100 W) for agarose melting
11. Vacuum filtration system (Nalgene, catalog number: 300-4050)
Software and datasets
1. ImageJ (https://imagej.net/ij/download.html)
2. GraphPad Prism 9.1.1 (https://www.graphpad.com/features)
Procedure
文章信息
稿件历史记录
提交日期: Nov 27, 2025
接收日期: Feb 18, 2026
在线发布日期: Mar 4, 2026
出版日期: Mar 20, 2026
版权信息
© 2026 The Author(s); This is an open access article under the CC BY-NC license (https://creativecommons.org/licenses/by-nc/4.0/).
如何引用
Cervantes-Rivera, R., González-Fernández, L. N., Rosas, A. Z. R., Ortíz, S. J. F., Ochoa-Zarzosa, A. and López-Meza, J. E. (2026). A Simple and Cost-Effective Method for Generating Spheroids From Triple-Negative Breast Cancer Cell Line (MDA-MB-231). Bio-protocol 16(6): e5641. DOI: 10.21769/BioProtoc.5641.
分类
癌症生物学 > 通用技术 > 细胞生物学试验
干细胞 > 类器官培养
细胞生物学 > 细胞分离和培养 > 3D细胞培养
您对这篇实验方法有问题吗?
在此处发布您的问题,我们将邀请本文作者来回答。同时,我们会将您的问题发布到Bio-protocol Exchange,以便寻求社区成员的帮助。
Share
Bluesky
X
Copy link