- EN - English
- CN - 中文
A Cytosine Deaminase–Based Genomic Footprinting Assay (cFOOT-seq) for Detecting Transcription Factor Occupancy
(*contributed equally to this work) 发布: 2026年03月20日第16卷第6期 DOI: 10.21769/BioProtoc.5637 浏览次数: 21
评审: Alba BlesaGuohao HanAnonymous reviewer(s)

相关实验方案

HaloChIP-seq用于体内小鼠转录因子 Cistromes 的抗体独立映射
Ann Louise Hunter [...] David A. Bechtold
2022年07月05日 4104 阅读
用于在活细胞中以单个核苷酸分辨率分析RNA-蛋白质相互作用位点的修订iCLIP-seq方案
Syed Nabeel-Shah and Jack F. Greenblatt
2023年06月05日 5236 阅读
Abstract
Transcription factors (TFs) regulate gene expression by binding to cis-regulatory elements in the genome. Understanding transcriptional regulation requires genome-wide characterization of TF occupancy across different chromatin contexts, yet simultaneous assessment of TF binding for multiple factors remains technically challenging. Here, we describe a detailed and reproducible protocol for cFOOT-seq, a cytosine deaminase–based genomic footprinting assay by sequencing, which enables antibody-independent, base-resolution profiling of chromatin accessibility, nucleosome organization, and TF occupancy. In cFOOT-seq, the double-stranded DNA (dsDNA) cytosine deaminase SsdAtox converts cytosine to uracil in accessible chromatin, whereas TF binding and nucleosome occupancy locally protect DNA from deamination. Using the FootTrack analysis framework, deamination patterns generated by cFOOT-seq are quantitatively analyzed to derive standardized footprint and chromatin organization profiles at base resolution across the genome. Because cFOOT-seq preserves genomic DNA integrity during deamination-based footprinting, it is compatible with ATAC-seq-based chromatin enrichment. ATAC-combined implementations reduce sequencing depth requirements and improve scalability for footprint-focused analyses, supporting applications in low-input and single-cell settings. This protocol provides a practical framework for genome-wide TF footprint profiling and can be readily applied to dissect gene regulatory mechanisms in development, immunity, and disease, including cancer.
Key features
• cFOOT-seq provides antibody-independent, base-resolution, genome-wide profiling of TF occupancy together with chromatin accessibility and nucleosome organization in a single assay.
• Using the FootTrack analysis framework, cFOOT-seq enables quantitative and comparative analysis of TF footprint patterns and occupancy across the genome.
• cFOOT-seq preserves genomic DNA integrity during deamination-based footprinting and is compatible with ATAC-seq-based chromatin enrichment.
• Integrated with ATAC, cFOOT-seq enables sensitive and in-depth TF footprint analyses at low sequencing cost, supporting single-cell applications.
Keywords: Gene transcriptionGraphical overview
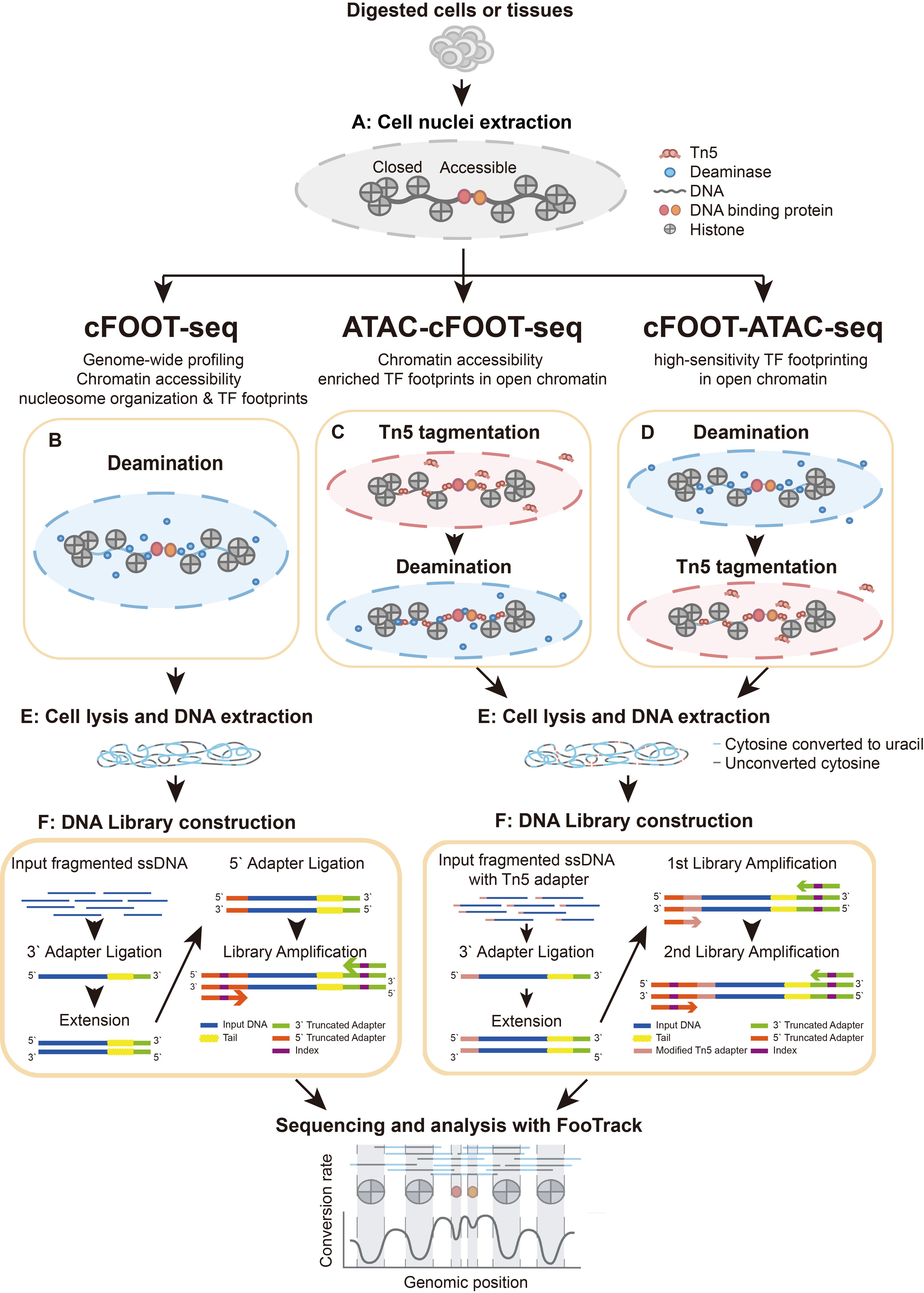
Overview of cFOOT-seq, ATAC-cFOOT-seq, and cFOOT-ATAC-seq procedure
Background
Gene regulation is often mediated by the coordinated activity of multiple transcription factors (TFs) acting at cis-regulatory elements across the genome [1,2]. At cis-regulatory elements, TF occupancy provides a direct readout of where regulatory interactions occur on chromatin. Understanding transcriptional regulation therefore requires approaches that can capture genome-wide TF occupancy information for many TFs in a comparable and integrated manner [3]. However, experimental strategies that enable such large-scale TF occupancy profiling remain limited.
Antibody-based methods for profiling TF binding, including ChIP-seq [4], CUT&RUN [5], CUT&Tag [6–9], DiMeLo-seq [10], and nanoHiMe-seq [11], directly measure protein–DNA interactions in chromatin but depend on antibody availability and specificity and typically profile one TF at a time. This limits scalability and makes it difficult to compare occupancy patterns across multiple TFs within the same sample. TF occupancy can also be inferred computationally from chromatin accessibility data generated by DNase-seq [12,13] or ATAC-seq [14,15]. Although these footprinting approaches are antibody-independent, their sensitivity is influenced by enzyme sequence bias, sequencing depth, and local cutting frequency, and they rely on population-averaged signals rather than directly encoding TF occupancy at the level of individual DNA molecules [16–18].
Single-molecule enzymatic footprinting strategies overcome these limitations by recording protein–DNA interactions as protection patterns against enzymatic modification of DNA. These include methyltransferase-based methods (e.g., NOMe-seq [19], SMF [20], SMAC-seq [21], and Fiber-seq [22]), as well as a growing number of recently reported approaches based on double-stranded DNA (dsDNA) cytosine deaminases, including FOODIE [23], cFOOT-seq [24], TDAC-seq [25], DAF-seq [26], and ACCESS-ATAC [27]. In deaminase-based footprinting, cytosine bases in accessible DNA are converted to uracil, whereas DNA regions occupied by TFs or nucleosomes are protected from deamination. After PCR amplification and sequencing, these protection patterns can be decoded as base-resolution footprints.
cFOOT-seq (a cytosine deaminase-based genomic footprinting assay by sequencing) implements this strategy using SsdAtox [28], a dsDNA cytosine deaminase with robust activity, low sequence bias, and minimal sensitivity to cytosine methylation [24]. This enables antibody-independent, base-resolution profiling of chromatin accessibility, nucleosome organization, and TF occupancy across the genome. Because cFOOT-seq preserves genomic DNA integrity during deamination-based footprinting, it can be combined with ATAC-seq. ATAC-integrated cFOOT-seq workflows enable footprint-focused analyses with reduced sequencing depth and support applications in low-input and single-cell settings. This protocol describes the experimental procedures and the FootTrack-based data analysis framework for cFOOT-seq and its ATAC-integrated workflows.
Materials and reagents
Biological materials
1. K562 (Procell, catalog number: CL-0130)
2. HepG2 (Procell, catalog number: CL0103)
Reagents
1. 1 M Tris-HCl, pH 7.4 (Bioworld, catalog number: 21420063-1)
2. 1 M Tris-HCl, pH 7.5 (Thermo Scientific, catalog number: 15567027)
3. 5 M sodium chloride (NaCl) (Sigma-Aldrich, catalog number: S5150)
4. 0.5 M ethylene diamine tetraacetic acid (EDTA) (Thermo Scientific, catalog number: 15575020)
5. Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: B2064)
6. 10% Tween-20 (Sigma-Aldrich, catalog number: 655206-50ML)
7. 10% NP-40 (Thermo Scientific, catalog number: 85124)
8. 5% digitonin (Thermo Scientific, catalog number: BN2006)
9. 1 M magnesium chloride (MgCl2) (Sigma-Aldrich, catalog number: 63069)
10. COmpleteTM EDTA-free protease inhibitor cocktail (PIC) (Sigma-Aldrich, catalog number: 04693132001)
11. Dimethylformamide (DMF) (Sigma-Aldrich, catalog number: 227056)
12. Ultrapure DNase/RNase-free distilled water (Thermo Scientific, catalog number: 10977023)
13. Dulbecco's phosphate-buffered saline (DPBS) (Gibco, catalog number: C14190500CP)
14. SsdAtox (DNA cytosine deaminase) (in-house purified; aliquot and store at -80 °C)
15. Uracil glycosylase inhibitor (UGI) (New England Biolabs, catalog number: M0281L)
16. Tn5 transposase (tagmentation enzyme; example used: TruePrep® Tagment Enzyme kit, Vazyme, catalog number: S111)
Note: Equivalent Tn5 enzymes from other suppliers can be used. Follow the manufacturer’s instructions for adapter assembly, and a small-scale test is recommended when switching reagents.
17. Lambda DNA (Thermo Scientific, catalog number: SD0021)
18. Glycogen (Sigma-Aldrich, catalog number: 10901393001)
19. Phenol:chloroform:isoamyl alcohol 25:24:1, pH ≥ 7.8 (PCI) (JSENB, catalog number: JS0700-100ML)
20. 100% ethanol (Sinopharm Chemical Reagent, catalog number: 10009218)
21. Isopropanol (Sinopharm Chemical Reagent, catalog number: 40064360)
22. Sodium bicarbonate (NaHCO3) (Sigma-Aldrich, catalog number: S5761-500G)
23. 3 M Sodium acetate buffer solution (NaAc) (Sigma-Aldrich, catalog number: S7899)
24. QubitTM ssDNA Assay kit (Thermo Scientific, catalog number: Q10212)
25. QubitTM dsDNA HS Assay kit (Thermo Scientific, catalog number: Q32854)
26. ssDNA Methylation-Style Library Preparation kit (example used: EpiArt DNA Methylation Library kit for Illumina V3, Vazyme, catalog number: NE103)
Note: Other kits with the same functionality (ssDNA library construction and compatibility with deaminated DNA templates) can also be used. A small-scale test is recommended to confirm library yield and fragment-size distribution.
27. DNA clean-up magnetic beads (SPRI-based) (example used: VAHTS DNA Clean Beads, Vazyme, catalog number: N411)
Note: Equivalent SPRI beads can be substituted. A small-scale test is recommended to confirm recovery and size selection performance.
28. Oligonucleotides for Tn5 transposome generation and library preparation (listed in Table 1)
Table 1. Sequences of oligonucleotides used in this protocol
| Name | Sequence (5′→3′) | Purpose | Notes/source |
|---|---|---|---|
| Tn5 primer D | TGGTAGAGAGGGTGAGATGTGTATAAGAGACAG | Cytosine-free Tn5 adapter (top strand) | Custom oligo; lacks cytosines |
| Tn5 ME oligo | phos-CTGTCTCTTATACACATCT-NH2 | Universal Tn5 mosaic end | Modified 5′ phosphate, 3′ NH2 |
| i5_bridge primer | ACACTCTTTCCCTACACGACGCTCTTCCGATCTTGGTAGAGAGGGTGAGATGTGTATAAGAGACAG | Add 5′ sequencing handle during pre-amplification | Custom oligo |
| Universal i7 primer | Provided in index primer kit | PCR amplification and indexing | Illumina-compatible i7 index primer |
| Universal i5 primer | Provided in index primer kit | PCR amplification and indexing | Illumina-compatible i5 index primer |
Solutions
1. 10% BSA in H2O (see Recipes)
2. 10% BSA in DPBS (see Recipes)
3. Transposome generation (see Recipes)
4. Nuclei extraction buffer (see Recipes)
5. Stop buffer (see Recipes)
6. 2× SsdAtox reaction buffer (2× SRB) (see Recipes)
7. SsdAtox-UGI mix (see Recipes)
8. Cell suspension buffer (see Recipes)
9. Hot lysis buffer (see Recipes)
10. 2× TD buffer (see Recipes)
11. Transposition reaction mix (see Recipes)
12. Tn5 washing buffer (see Recipes)
13. ATAC-cFOOT-cell suspension buffer (see Recipes)
14. 2× SsdAtox reaction buffer plus DPBS and BSA (2× SRB+DB) (see Recipes)
15. cFOOT-ATAC-SsdAtox-UGI mix (see Recipes)
16. cFOOT-ATAC-cell suspension buffer (see Recipes)
17. Deaminase washing buffer (see Recipes)
Recipes
1. 10%BSA in H2O
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Ultrapure DNase/RNase-free distilled water | N/A | Up to 10 mL |
| BSA powder | 10% (w/v) | 1 g |
| Total | N/A | 10 mL |
Before use, filter the solution through the Millex®-HV filter unit. This buffer should be stored at -20 °C.
2. 10% BSA in DPBS
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| DPBS | N/A | Up to 10 mL |
| BSA powder | 10% (w/v) | 1 g |
| Total | N/A | 10 mL |
Before use, filter the solution through the Millex®-HV filter unit. This buffer should be stored at -20 °C.
3. Transposome generation
Transposome assembly was performed using a Tn5-based workflow adapted from standard tagmentation procedures (example used: TruePrep® Tagment Enzyme kit, Vazyme). Tn5 transposase from other suppliers can also be used. However, the cytosine-free adapters described below should be used to avoid cytosine deamination by SsdAtox.
The adapter was generated by annealing Tn5 Primer D (lacking cytosines in its 5′ overhang) with the universal Tn5 ME oligo. The annealed adapter was then loaded onto Tn5 transposase to form the active transposome, following the manufacturer’s recommended annealing and loading instructions for the chosen Tn5 system. The assembled transposome can be aliquoted and stored at -20 °C (avoid repeated freeze–thaw cycles).
4. Nuclei extraction buffer
| Reagent | Final concentration | Volume for 10 reactions |
|---|---|---|
| 1 M Tris-HCl, pH 7.4 | 10 mM | 5 μL |
| 5 M NaCl | 10 mM | 1 μL |
| 0.1 M EDTA | 0.1 mM | 0.5 μL |
| 10% BSA in H2O | 1% | 50 μL |
| 10% Tween-20 | 0.1% | 5 μL |
| 10% NP-40 | 0.1% | 5 μL |
| 5% Digitonin | 0.01% | 1 μL |
| 1 M MgCl2 | 3 mM | 1.5 μL |
| 50× PIC | 1× | 10 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | 421 μL |
| Total | N/A | 500 μL |
Prepare immediately before use and keep at 0–4 °C until use. The commercially available EDTA solution is 0.5 M and should be diluted to 0.1 M for use as a stock solution when preparing the nuclei extraction buffer. Optional: The nuclei extraction buffer without PIC and any detergents can be prepared ahead of time and stored at -20 °C for up to 1 month. Add the detergents and PIC immediately before use.
5. Stop buffer
| Reagent | Final concentration | Volume for 10 reactions |
|---|---|---|
| 1 M Tris-HCl, pH 7.4 | 10 mM | 100 μL |
| 5 M NaCl | 10 mM | 20 μL |
| 0.1 M EDTA | 0.1 mM | 10 μL |
| 10% BSA in H2O | 1% | 1 mL |
| Ultrapure DNase/RNase-free distilled water | N/A | 8870 μL |
| Total | N/A | 10 mL |
Prepare immediately before use and keep at 0–4 °C until use.
6. 2× SsdAtox reaction buffer (2× SRB)
| Reagent | Final concentration | Volume for 10 reactions |
|---|---|---|
| 1 M Tris-HCl, pH 7.4 | 20 mM | 5 μL |
| 0.1 M DTT | 2 mM | 5 μL |
| 50× PIC | 2× | 10 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | 230 μL |
| Total | N/A | 250 μL |
Prepare immediately before use and keep at 0–4 °C until use.
7. SsdAtox-UGI mix
| Reagent | Final concentration | Volume per reaction (1×) |
|---|---|---|
| 2× SRB | 1× | 10 μL |
| 2 U/μL UGI | 0.1 U/μL | 2.5 μL |
| SsdAtox | 3–20 U/μL | × μL |
| Ultrapure DNase/RNase-free distilled water | N/A | Up to 20 μL |
Prepare immediately before use and keep at 0–4 °C until use.
8. Cell suspension buffer
| Reagent | Final concentration | Volume per reaction (1×) |
|---|---|---|
| 2× SRB | 1× | 15 μL |
| 0.4 ng/μL Lambda DNA | 0.04 ng/μL | 5 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | Up to 30 μL |
Prepare immediately before use and keep at 0–4 °C until use.
9. Hot lysis buffer
| Reagent | Final concentration | Volume per reaction (1×) |
|---|---|---|
| 1 M NaHCO3 | 100 mM | 25 μL |
| 10% SDS | 1% | 5 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | 150 μL |
| Total | N/A | 200 μL |
Prepare immediately before use and keep at room temperature.
10. 2× TD buffer
| Reagent | Final concentration | Volume per reaction (1×) |
|---|---|---|
| 1 M Tris-HCl, pH 7.5 | 20 mM | 0.5 μL |
| 1 M MgCl2 | 10 mM | 0.25 μL |
| 100% DMF | 20% | 5 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | 19.25 μL |
| Total | N/A | 25 μL |
Prepare immediately before use and keep at 0–4 °C until use.
11. Transposition reaction mix
| Reagent | Final concentration | Volume per reaction (1×) |
|---|---|---|
| 2× TD buffer | 1× | 25 μL |
| DPBS | 33% (v/v) | 16.5 μL |
| Transposase | 100 nM | 0.667 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | 7.833 μL |
| Total | N/A | 50 μL |
Prepare fresh on ice immediately before use and keep at 0–4 °C until use.
12. Tn5 washing buffer
| Reagent | Final concentration | Volume for 1× reaction |
|---|---|---|
| 2× SRB | 1× | 250 μL |
| 0.1 M EDTA | 454.5 nM | 2.5 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | 247.5 μL |
| Total | N/A | 500 μL |
Prepare immediately before use and keep at 0–4 °C until use.
13. ATAC-cFOOT-cell suspension buffer
| Reagent | Final concentration | Volume for 1× reaction |
|---|---|---|
| 2× SRB | 1× | 15 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | 15 μL |
| Total | N/A | 30 μL |
Prepare immediately before use and keep at 0–4 °C until use.
14. 2× SsdAtox reaction buffer plus DPBS and BSA (2× SRB+DB)
| Reagent | Final concentration | Volume for 10 reactions |
|---|---|---|
| 1 M Tris-HCl, pH 7.4 | 20 mM | 5 μL |
| 0.1 M DTT | 2 mM | 5 μL |
| DPBS | 66% (v/v) | 165 μL |
| 10% BSA in H2O | 0.04% | 1 μL |
| 50× PIC | 2× | 10 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | 64 μL |
| Total | N/A | 250 μL |
Note: We typically prepare 2× SRB+DB fresh (with DTT and PIC) and keep it at 0–4 °C until use. Optional: A 2× SRB+DB base without DTT and PIC can be prepared in advance, aliquoted, and stored at -20 °C for up to 1 month. Add DTT and PIC immediately before use.
15. cFOOT-ATAC-SsdAtox -UGI mix
| Reagent | Final concentration | Volume per reaction (1×) |
|---|---|---|
| 2× SRB+DB | 1× | 10 μL |
| 2 U/μL UGI | 0.1 U/μL | 2.5 μL |
| SsdAtox | N/A | × μL |
| Ultrapure DNase/RNase-free distilled water | N/A | Up to 20 μL |
Prepare immediately before use and keep at 0–4 °C until use.
16. cFOOT-ATAC-cell suspension buffer
| Reagent | Final concentration | Volume per reaction (1×) |
|---|---|---|
| 2× SRB+DB | 1× | 15 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | 15 μL |
| Total | N/A | 30 μL |
Prepare immediately before use and keep at 0–4 °C until use.
17. Deaminase washing buffer
| Reagent | Final concentration | Volume per reaction (1×) |
|---|---|---|
| 2× SRB+DB | 1× | 250 μL |
| Ultrapure DNase/RNase-free distilled water | N/A | 250 μL |
| Total | N/A | 500 μL |
Prepare fresh immediately before use and keep at 0–4 °C until use.
Note: Before the experiment, pre-mix well every buffer above.
Laboratory supplies
1. 1.5 mL DNase/RNase-free tubes (Axygen, catalog number: MCT-150-C)
2. 0.2 mL DNase/RNase-free PCR tubes (Axygen, catalog number: PCR-02-C)
3. 50 mL conical tubes (Biobest, catalog number: BSD-HCT-50A)
4. 15 mL conical tubes (Biobest, catalog number: BSD-HCT-15A)
5. 1.5 mL DNA LoBind® tube (Eppendorf, catalog number: 0030108051)
6. 10 μL filter pipette tips (Biobest, catalog number: BSD-HTF-CTL-10)
7. 20 μL filter pipette tips (Biofil, catalog number: PMT252520)
8. 100 μL filter pipette tips (Biofil, catalog number: PMT252100)
9. 200 μL filter pipette tips (Biofil, catalog number: PMT231200)
10. 1 mL filter pipette tips (Biobest, catalog number: BSD-HTF-1250-AR)
11. Millex®-HV filter unit (Millipore, catalog number: SLHV033RB)
12. microTUBE-50 AFA fiber screw-cap case (Covaris, catalog number: 520166)
Equipment
1. FinnpipetteTM F1 variable volume single-channel pipettes, 100–1,000 μL (Thermo Scientific, catalog number: 4641100N)
2. FinnpipetteTM F1 variable volume pipettes, 20–200 μL (Thermo Scientific, catalog number: 4641080N)
3. FinnpipetteTM F1 variable volume pipettes, 10–100 μL (Thermo Scientific, catalog number: 4641070N)
4. FinnpipetteTM F1 variable volume pipettes, 2–20 μL (Thermo Scientific, catalog number: 4641060N)
5. FinnpipetteTM F1 variable volume pipettes, 1–10 μL (Thermo Scientific, catalog number: 4641030N)
6. FinnpipetteTM F1 variable volume pipettes, 0.2–2 μL (Thermo Scientific, catalog number: 4641010N)
7. Thermomixer (Yeasen, catalog number: 80440ES03)
8. QubitTM 4 NGS Starter Kit, with WiFi (Thermo Scientific, catalog number: Q33240)
9. Refrigerated centrifuge (Eppendorf, model: 5425 R)
10. M220 Focused ultrasonicator (Covaris, catalog number: 500295)
11. T100 thermal cycler (Bio-Rad, catalog number: 1861096)
Software and datasets
1. FastQC (v0.12.1): Quality control of raw sequencing reads, including assessment of base quality scores, GC content, sequence duplication levels, and overrepresented sequences. Source: https://www.bioinformatics.babraham.ac.uk/projects/fastqc
2. Trim Galore (v0.6.10): Adapter trimming and removal of low-quality bases from sequencing reads. Source: https://www.bioinformatics.babraham.ac.uk/projects/trim_galore
3. BASAL (v1.8): Conversion-aware mapping of nucleotide base-conversion sequencing reads. Source: https://github.com/JiejunShi/BASAL
4. BasalKit (v1.8): Extraction of per-base sequencing coverage and C-to-T conversion rates from BASAL-aligned BAM files, followed by generation of base-conversion tables for downstream analysis. BasalKit is an integrated utility within the BASAL framework specifically designed for base-conversion sequencing data. Source: https://github.com/JiejunShi/BASAL (distributed as part of the BASAL package)
5. bedtools (v2.31.1): Removal of genomic positions overlapping known single-nucleotide polymorphisms (SNPs) to prevent interference with the accurate estimation of base-conversion rates. Source: https://bedtools.readthedocs.io/en/latest
6. FootTrack (v1.1.0): Comprehensive analysis of transcription factor occupancy from conversion-based sequencing data, including bias correction, footprint scoring, differential TF binding analysis, and de novo TF binding site prediction. Source: https://github.com/ZhangLab-TJ/FootTrack
7. ATAC-seq and transcription factor ChIP-seq datasets used in this protocol were primarily obtained from the ENCODE Project and the Cistrome Project (https://www.encodeproject.org; http://cistrome.org). Detailed descriptions of the datasets, including experiment identifiers and accession numbers, can be found in the original article [24]. Transcription factor motif files were downloaded from the JASPAR database (https://jaspar.genereg.net) and used for genome-wide motif scanning and footprint-based TF binding analysis.
Single-nucleotide polymorphism data for mouse genomes were obtained from the Mouse Genomes Project (https://www.mousegenomes.org/snps-indels). SNP information for human cell lines was retrieved from the ENCODE Project (https://www.encodeproject.or/) and used to mask polymorphic positions during base-conversion rate calculations.
Procedure
文章信息
稿件历史记录
提交日期: Dec 23, 2025
接收日期: Feb 11, 2026
在线发布日期: Feb 27, 2026
出版日期: Mar 20, 2026
版权信息
© 2026 The Author(s); This is an open access article under the CC BY-NC license (https://creativecommons.org/licenses/by-nc/4.0/).
如何引用
Yang, M. C., Wu, A., Wang, H., Liu, X., Shi, J., Gao, S. and Zhang, J. M. (2026). A Cytosine Deaminase–Based Genomic Footprinting Assay (cFOOT-seq) for Detecting Transcription Factor Occupancy. Bio-protocol 16(6): e5637. DOI: 10.21769/BioProtoc.5637.
分类
发育生物学 > 细胞信号传导
分子生物学 > DNA > DNA-蛋白质相互作用
您对这篇实验方法有问题吗?
在此处发布您的问题,我们将邀请本文作者来回答。同时,我们会将您的问题发布到Bio-protocol Exchange,以便寻求社区成员的帮助。
Share
Bluesky
X
Copy link