- EN - English
- CN - 中文
Fluorescence-Based Absent Allele-Specific Amplification (FAASA) for High-Throughput Detection of Absent Alleles
(*contributed equally to this work) 发布: 2026年03月20日第16卷第6期 DOI: 10.21769/BioProtoc.5633 浏览次数: 29
评审: Lucy XieKirsten A. CoprenAhana Byne
Abstract
In wheat and other crops, some genes display presence/absence variation, and it is occasionally beneficial to select for the absent allele to remove a functional gene. However, current high-throughput genotyping methods used to detect the absence of genes tend to be inconsistent and inconclusive. Kompetitive allele-specific PCR (KASP) and PCR allele competitive extension (PACE) are two well-established methods for allele-specific polymerase chain reaction (AS-PCR) assays, each using fluorescence resonance energy transfer (FRET) to generate a signal for each allele, typically targeting biallelic single-nucleotide polymorphisms. KASP and PACE methods are more difficult to apply to alleles with presence/absence variation because the lack of amplification of the absent allele is indistinguishable from a failed PCR. Here, we present a multiplex fluorescence-based absent allele–specific amplification (FAASA) method using the PACE marker system (compatible with KASP markers) to detect the absence of one particular or all alleles of a target sequence using a primer mix consisting of one target-specific primer pair (TSP) and a second primer set specific to a highly conserved endogenous gene known as a core gene–specific primer pair (CGSP). The forward primer of each pair is tagged with a 5′ terminal tail complementary to dye-labeled oligonucleotides in commercially available FRET cassettes. Lines that amplify only the core gene do not carry the target, while lines that amplify both the core gene and the target carry alleles of both the core gene and the target. The inclusion of the CGSPs allows researchers to confidently distinguish lines with absent alleles of the target from lines with failed PCR reactions, which can happen due to various reasons, including inadequate DNA quality or quantity.
Key features
• A robust, affordable, high-throughput genotyping method for genes or other target sequences with presence/absence variation.
• FAASA markers can be easily incorporated into established marker-assisted selection programs in labs using KASP and/or PACE markers.
• FAASA markers can also be used for other genotyping applications like GWAS, QTL, or bi-parental mapping studies.
• Easily adaptable to different targets and species of interest.
Keywords: GenotypingGraphical overview
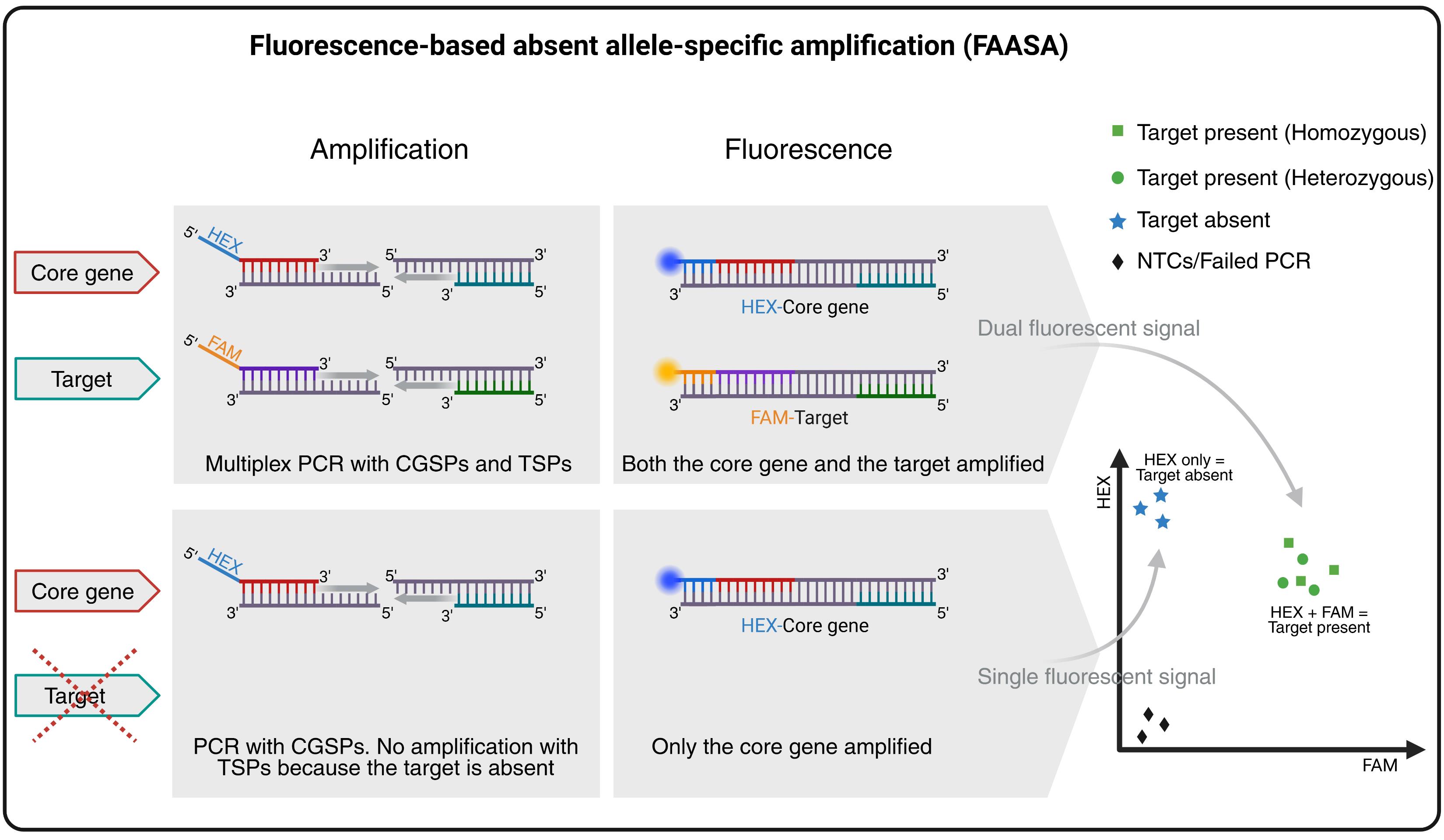
Overview of the fluorescence-based absent allele–specific amplification (FAASA) methodology. Created with BioRender.com.
Background
In wheat, there are certain genes, such as disease susceptibility genes, that need to be eliminated to develop favorable varieties. However, developing a diagnostic high-throughput marker for marker-assisted selection of genes that show presence/absence variation is challenging. The absence of the target gene results in no amplification during PCR, which makes it impossible to distinguish a sample with a truly absent allele from one with a failed PCR, which can occur due to technical issues. Here, we present a high-throughput marker system named fluorescence-based absent allele–specific amplification (FAASA) to confidently detect absent alleles of any target of interest (gene, a particular allele of a gene, promoter, indel, etc.).
This approach is based on multiplex PCR, a technique first described by Chamberlain et al. in 1988 [1] that has since been used in various assays [2–4]. In general, multiplex PCR refers to the amplification of multiple amplicons simultaneously during PCR using more than two primers in the same reaction mixture. It is commonly used in allele-specific PCR assays such as kompetitive allele-specific PCR (KASP) and PCR allele competitive extension (PACE) genotyping systems [5], where two amplicons are amplified based on a single-nucleotide polymorphism (SNP) between two alleles of the same target sequence. Adaptations of the original KASP/PACE systems to amplify two target regions, rather than SNP variations, have also been reported [6–8]. In FAASA, we utilize the PACE system to simultaneously amplify two distinct genomic regions (target and core gene) to enable high-throughput detection of absent alleles. Here, a total of four primers are used in a multiplex PCR: one pair of target-specific primers (TSPs) and one pair of core gene–specific primers (CGSPs). The forward primers of both TSPs and CGSPs are tagged with FAM and HEX (fluorescein amidite and hexachlorofluorescein) fluorescent tails, respectively. The core gene is expected to be present in all biological samples. As a result, the CGSPs will consistently amplify a fragment of the core gene, producing a HEX fluorescent signal. This signal serves as an internal control, indicating that the PCR reaction was successful. Simultaneously, the TSPs will amplify a fragment of the target, generating a FAM signal if the target is present in the sample. As a result, samples containing the target will produce both FAM and HEX signals. These overlapping signals create a combined fluorescence pattern that typically appears as a cluster in the middle of the endpoint fluorescence scatter plot, resembling a heterozygous cluster. However, this is not due to actual heterozygosity but rather to the simultaneous detection of both amplified core gene and target products. If the target is absent from a sample, only the core gene will be amplified, resulting in a HEX-only signal. These samples will form a separate cluster corresponding to the absence of the target. Samples with failed PCR reactions and non-template controls (NTCs) will show no amplification for either of the genes and will cluster in the bottom-left corner of the scatter plot due to the absence of both signals. While we report the use of FAM for the target and HEX for the core gene, any two compatible fluorescent dyes may be used, and their assignment may also be switched if preferred.
While this method can reliably detect the absence of a target, it is important to note that it does not distinguish between homozygous and heterozygous states when the target is present. However, FAASA can confidently and efficiently differentiate truly absent alleles from PCR failure, which can be particularly valuable for plant breeders during large-scale marker-assisted selections, often allowing the bypass of exhaustive phenotypic screens. While FAASA markers can be designed to detect the absence of a particular allele or all alleles of a target sequence, they cannot account for accrued mutations within the target that are not targeted by the TSPs that potentially render the target nonfunctional. Thus, if there are multiple segregating mutations in a target that produce the same phenotype, multiple FAASA markers may be needed to adequately detect all samples with alleles producing that phenotype. This method can be applied more broadly to detect any target in a genome, genic or non-genic, as long as primers can be designed to specifically amplify the target of interest. Therefore, these markers can also be used in molecular mapping projects. While we present an example in this protocol using a FAASA marker that amplifies wheat target and core genes, FAASA markers may be used in other species and scenarios as well.
Materials and reagents
Biological materials
1. 45 ng of dsDNA or 90 ng of total DNA per biological sample per replicate; include at least one control sample with the target as a positive control and a sample without the target as a negative control. Appropriate controls are described in more detail in section A.
Note: This quantity of DNA is specific to wheat, which has a very large genome. If applying this method to a species with a smaller genome, less DNA may be needed, as the relative concentration of the target and core genes will be higher in a smaller genome.
Reagents
1. ddH2O
2. One pair of target-specific primers (TSPs) (see section A)
a. 100 μM FAM tagged TSP (5′ FAM terminal tail “GAAGGTGACCAAGTTCATGCT”)
b. 100 μM reverse TSP
c. The following target-specific primers for the FAASA marker fcp1069 were used in this protocol as an example: Tsc1-1Ka-Null-Fam: GAAGGTGACCAAGTTCATGCTATAAACGAAAGATACTTGTTTTGCTC, Tsc1-1Ka-Null-Rev: ATTATTATGACAGAAATTGCAACAACA
3. One pair of core gene–specific primers (CGSPs) (see section A)
a. 100 μM HEX tagged CGSP (5′ HEX terminal tail “GAAGGTCGGAGTCAACGGATT”)
b. 100 μM reverse CGSP
c. The following core gene–specific primers were used for the marker fcp1069 in this protocol: Q_KB.1-Hex: GAAGGTCGGAGTCAACGGATTGCTAATTAAACGTCCACAGCAT, Q_KR.1-Rev: GACACTAATTAGTAGTAGATGTGACAG
Note: The sequences provided for the TSPs and CGSPs are for marker fcp1069, targeting the wheat genes Tsc1 and Q gene, respectively. Marker fcp1069 can be used to detect the absence of the tan spot susceptibility gene Tsc1 in wheat. The CGSPs can be used in conjunction with other TSPs to create FAASA markers to detect the absence of other targets in wheat.
4. 2× PACE genotyping master mix with standard ROX concentration (3CR Bioscience, catalog numbers: 001-0001, 001-0002, 001-0003, or 001-0004)
Note: Fluorescent probes in the master mix are light sensitive. To minimize exposure to light and prevent degradation from repeated freeze-thaw cycles, freeze small aliquots at -20 °C. Thaw 2× PACE genotyping master mix in a drawer or under cover. There are commercially produced light-blocking microcentrifuge tubes that may be a better option if light exposure cannot be adequately controlled. Use the FAASA master mix immediately.
5. 70% (v/v) ethanol
Solutions
1. FAASA primer mix (see Recipes)
2. FAASA master mix (see Recipes)
Recipes
1. FAASA primer mix
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| 100 μM FAM tagged TSP (Tsc1-1Ka-Null-Fam) | 12 μM | 12 μL |
| 100 μM reverse TSP (Tsc1-1Ka-Null-Rev) | 30 μM | 30 μL |
| 100 μM HEX tagged CGSP (Q_KB.1-Hex) | 12 μM | 12 μL |
| 100 μM reverse CGSP (Q_KR.1-Rev) | 30 μM | 30 μL |
| ddH2O | 16 μL | |
| Total | 73× | 100 μL |
Note: The primers in parentheses are the primers for marker fcp1069 used as an example in this protocol. Other primers may be used. Store at -20 °C and bring to room temperature before use.
2. FAASA master mix
| Reagent | Final concentration | Volume per reaction | Volume for 16 reactions | Volume for 96 reactions | Volume for 384 reactions |
|---|---|---|---|---|---|
| FAASA primer mix | 1× | 0.055 μL | 1.375 μL | 6.05 μL | 23.10 μL |
| 2× PACE genotyping master mix | 1× | 2 μL | 50 μL | 220 μL | 840 μL |
| ddH2O | 2 μL | 50 μL | 220 μL | 840 μL | |
| Total | 4.055 μL | 101.375 μL | 446.05 μL | 1,703.1 μL | |
| Volume per well in 8 wells | 11 μL (2.75 reactions) | 54 μL (13.5 reactions) | 210 μL (52.5 reactions) |
Notes:
1. Volumes for 16, 96, and 384 reactions include extra to account for volume lost during pipetting due to the saponaceous quality of 2× PACE genotyping master mix.
2. It is very difficult to see if there is liquid already pipetted into a well because the wells are opaque. We recommend dividing the master mix into 8 wells in a column on a PCR plate or strip and using a multichannel pipette to distribute the FAASA master mix to the PCR plate to reduce the chance of pipetting errors. The volume to put in each well is provided in the recipe.
Laboratory supplies
1. 20, 250, and 1,000 μL pipette tips (Rainin, catalog numbers: 17005872, 17005874, and 17007089, or similar)
2. (Optional) Filtered 20, 200, and 1,000 μL pipette tips (Rainin, catalog numbers: 30389227, 30389239, and 30389212, or similar)
3. 2, 20, and 1000 μL pipette (Rainin, catalog numbers: 17014393, 17014392, 17014382, or similar)
4. 10 μL multichannel pipette (Rainin, catalog number: 17013802)
5. 96-well PCR plate (BrandTech, catalog number: 781368, or similar)
6. (Optional) 8-well PCR strip tubes (BrandTech, catalog number: 781326, or similar)
7. 384-well PCR plate (Bio-Rad, catalog number: HSP3805, or similar)
8. TempPlate RT optical film (USA Scientific, catalog number: 2978-2100)
9. (Optional) AxySeal sealing film (Axygen, catalog number: PCR-SP, or similar)
10. MicroAmpTM adhesive film applicator (Thermo Fisher Scientific, catalog number: 4333183, or similar)
11. 1.7 or 2 mL microcentrifuge tubes, depending on master mix size (Fisherbrand, catalog numbers: 02-681-331 or 02-681-332, or similar)
Equipment
1. Real-time PCR machine or thermocycler and fluorescent plate reader (Bio-Rad CFX Opus 384 Real-Time PCR System, model: 12011452)
2. Spectrophotometer (Nanodrop, model: ND 8000)
3. Vortex (Fisherbrand, catalog number: 02-215-414)
4. Plate mixer (Eppendorf MixMate, catalog number: 5353000529)
5. Centrifuge with plate adapters (Eppendorf, model: Centrifuge 5810 R)
6. Incubator (Fisherbrand, catalog number: 15-103-0513, or similar)
Software and datasets
1. CFX Maestro (license required, comes with Bio-Rad CFX Opus 384 Real-Time PCR System), Bio-Rad, versions 1.1–2.3, or similar clustering software
2. (Optional) Primer3 (available online for free at https://www.ncbi.nlm.nih.gov/tools/primer-blast/)
Procedure
文章信息
稿件历史记录
提交日期: Dec 10, 2025
接收日期: Feb 5, 2026
在线发布日期: Feb 26, 2026
出版日期: Mar 20, 2026
版权信息
© 2026 The Author(s); This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/).
如何引用
Running, K. L. D., Seneviratne, S., Zhang, Z., Singh, G., Fiedler, J. D. and Faris, J. D. (2026). Fluorescence-Based Absent Allele-Specific Amplification (FAASA) for High-Throughput Detection of Absent Alleles. Bio-protocol 16(6): e5633. DOI: 10.21769/BioProtoc.5633.
分类
植物科学 > 植物育种
分子生物学 > DNA > 基因分型
您对这篇实验方法有问题吗?
在此处发布您的问题,我们将邀请本文作者来回答。同时,我们会将您的问题发布到Bio-protocol Exchange,以便寻求社区成员的帮助。
Share
Bluesky
X
Copy link