- EN - English
- CN - 中文
Simple Induction and Detection of Anthocyanins in Arabidopsis thaliana: A Tool for Mutant Screening and Metabolic Analysis
拟南芥花青素的简易诱导与检测方法:用于突变体筛选与代谢分析
发布: 2026年03月05日第16卷第5期 DOI: 10.21769/BioProtoc.5628 浏览次数: 38
评审: Shweta PanchalSujata KumariJose Roberto Torres
Abstract
Anthocyanins are specialized flavonoid pigments that play critical roles in plant coloration, photoprotection, and responses to environmental stress. Arabidopsis thaliana serves as a valuable genetic model for dissecting anthocyanin biosynthesis and regulatory networks. Conventional methods for anthocyanin quantification, such as crude spectrophotometric assays, often compromise pigment integrity, yield inconsistent results, and provide limited information on compound composition. Here, we describe a simple, reproducible, and high-fidelity protocol for the induction, extraction, quantification, and chromatographic profiling of anthocyanins in Arabidopsis thaliana seedlings. The workflow employs well-defined anthocyanin-inductive conditions (AIC), methanol/formic acid extraction, lyophilization for dry-weight normalization, and dual quantification via spectrophotometry and High-performance liquid chromatography with diode-array detection (HPLC-DAD) analysis. This protocol enables accurate comparison between wild-type and mutant genotypes, facilitating both mutant screening and metabolic pathway analysis. The approach minimizes pigment degradation, enhances reproducibility across replicates, and offers a robust tool for research in plant metabolism, stress physiology, and flavonoid biochemistry.
Key features
• This protocol establishes well-defined anthocyanin-inductive conditions (AIC) using sucrose and continuous light, enabling reproducible pigment accumulation in Arabidopsis thaliana seedlings.
• This protocol employs methanol/formic acid extraction and lyophilization to maintain anthocyanin stability and minimize degradation during sample processing.
• This protocol integrates spectrophotometric OD532 normalization with HPLC-DAD profiling for quantification of total anthocyanins and characterization of individual anthocyanin species.
• This protocol is suitable for mutant screening, metabolic pathway analysis, and stress-response studies in Arabidopsis thaliana.
Keywords: Anthocyanin-inductive conditions (AIC) (花青素诱导条件)Graphical overview
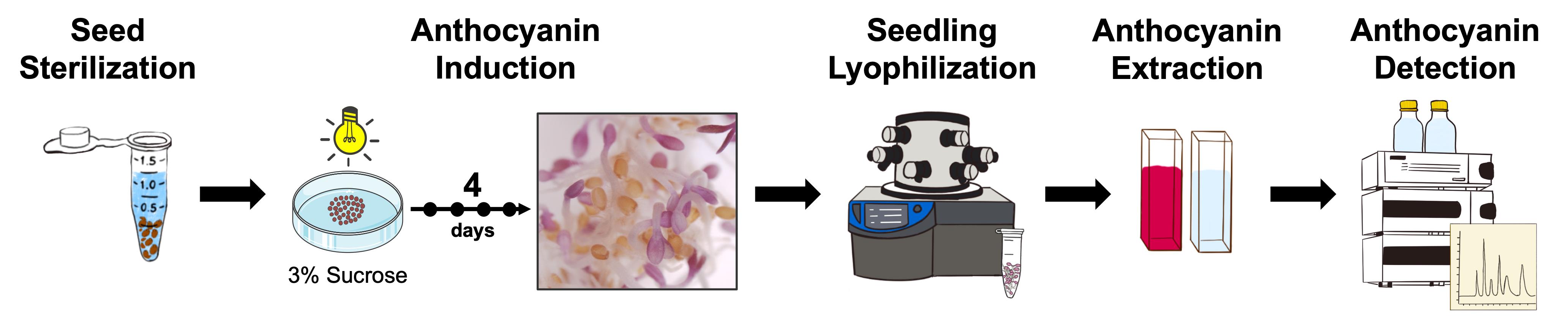
Graphical overview of anthocyanin induction, extraction, quantification, and HPLC-based detection in Arabidopsis thaliana. The schematic summarizes the complete workflow, including seed sterilization, anthocyanin induction under anthocyanin-inductive conditions (AIC), pigment extraction from lyophilized seedlings, spectrophotometric quantification, and high-performance liquid chromatography with diode-array detection (HPLC-DAD)-based detection of anthocyanins.
Background
Anthocyanins (Figure 1) are a class of water-soluble flavonoid pigments responsible for the red, purple, and blue coloration observed in many plant tissues [1,2]. Beyond their visual functions in attracting pollinators and serving as stress indicators [3,4], anthocyanins perform physiological roles by scavenging reactive oxygen species (ROS) and enhancing tolerance to abiotic stresses such as high light, nutrient deficiency, and low temperature [5,6]. Arabidopsis thaliana has become a widely used model system for investigating anthocyanin biosynthesis and regulation, owing to its fully sequenced genome, genetic tractability, and availability of numerous pigment-related mutants [7–9]. Its responsiveness to sucrose supplementation and continuous light exposure makes it particularly suitable for anthocyanin induction under controlled anthocyanin-inductive conditions (AIC) [10–14].
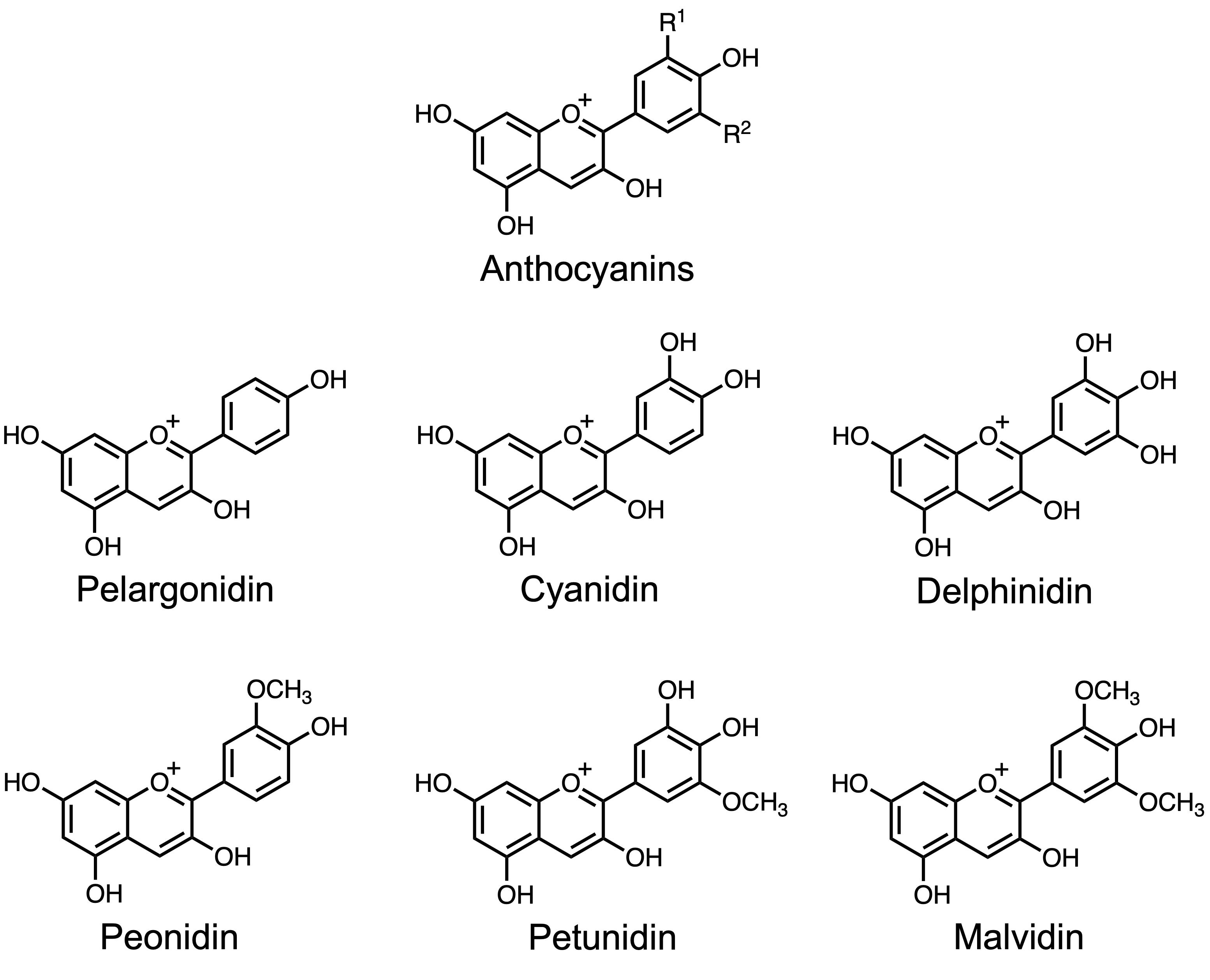
Figure 1. Chemical structures of representative anthocyanins. The basic anthocyanidin backbone consists of a flavylium cation structure with variable hydroxylation and methoxylation patterns that define specific anthocyanidins, such as pelargonidin, cyanidin, delphinidin, peonidin, petunidin, and malvidin. R1, R2: -H, -OH, -OCH3.
Traditional quantification of anthocyanins in Arabidopsis thaliana typically involves spectrophotometric assays of crude extracts, often based on absorbance at 530 nm with corrections for chlorophyll interference [15,16]. While these assays are rapid and cost-effective, they lack specificity in distinguishing individual anthocyanin species and provide limited resolution for comparative analyses among genotypes or treatments. In contrast, chromatographic techniques such as high-performance liquid chromatography (HPLC), coupled with diode array detection (DAD) or mass spectrometry [1], enable precise separation, identification, and quantification of individual anthocyanin compounds [12,17,18]. Several previously reported extraction methods rely on harsh conditions such as strong acid hydrolysis, which may result in modification of glycosylated forms [19,20].
The protocol presented here integrates optimized steps for anthocyanin induction, extraction, quantification, and chromatographic characterization in Arabidopsis thaliana. By combining a mild methanol/formic acid extraction system with lyophilization and normalization by dry weight, the method enhances pigment recovery, stability, and reproducibility. The inclusion of the chalcone isomerase mutant tt5–2 [9,21] alongside the wild-type Col-0 provides a robust system for comparing anthocyanin-deficient and anthocyanin-producing phenotypes, facilitating both genetic and metabolic analyses. Beyond Arabidopsis, this protocol can be adapted and optimized for other plant systems in which anthocyanin accumulation is environmentally or developmentally regulated, including crop and ornamental species. The protocol’s precision and reproducibility make it suitable for metabolic engineering and synthetic biology applications, where anthocyanins are targeted for enhancement of nutritional value, stress resilience, or natural pigment production in engineered plants and cell cultures.
Compared with existing methods, this protocol defines reproducible anthocyanin-inductive conditions (AIC), standardizes extraction buffers compatible with HPLC analysis, and integrates dual quantification approaches, spectrophotometric and chromatographic, for comprehensive pigment profiling. Beyond its utility in basic plant physiology, this approach is broadly applicable to studies of anthocyanin biosynthesis, stress physiology, metabolic engineering for pigment production, and functional genomics of the flavonoid pathway in Arabidopsis thaliana and other plant species [10,12,17,18].
Materials and reagents
Biological materials
1. Arabidopsis thaliana Col-0 seeds [Arabidopsis Biological Resource Center (ABRC), stock number: CS70000]
2. Arabidopsis thaliana tt5-2 seeds (ABRC, stock number: CS2105574)
Reagents
1. Ethanol (MiliporeSigma, catalog number: 64-17-5)
2. Bleach (Clorox, catalog number: USA001229)
3. Triton X-100 (VWR, catalog number: 9002-93-01)
4. Sucrose (FisherScientific, catalog number: 57-50-1)
5. Methanol (MeOH) (FisherScientific, catalog number: 67-56-1)
6. Formic acid (FisherScientific, catalog number: 270480010)
7. Acetonitrile (FisherScientific, catalog number: 75-05-08)
Solutions
1. 70% ethanol (see Recipes)
2. 30% bleach (see Recipes)
3. 0.1% Triton X-100 (see Recipes)
4. Sterilizing solution (see Recipes)
5. 3% sucrose (see Recipes)
6. Extraction buffer (see Recipes)
7. HPLC running buffer A (see Recipes)
8. HPLC running buffer B (see Recipes)
Recipes
1. 70% ethanol (500 mL)
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Ethanol | 70% (v/v) | 350 mL |
| dH2O | n/a | 150 mL |
| Total | n/a | 500 mL |
Store at room temperature (RT).
2. 30% bleach (500 mL)
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Commercial bleach | 30% (v/v) | 150 mL |
| dH2O | n/a | 350 mL |
| Total | n/a | 500 mL |
Store at RT.
3. 0.1% Triton X-100 (250 mL)
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Triton X-100 | 0.1% (v/v) | 0.25 mL |
| dH2O | n/a | 249.75 mL |
| Total | n/a | 250 mL |
Store at RT.
4. Sterilizing solution (50 mL)
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Commercial bleach | 27% (v/v) | 45 mL |
| Triton X-100 | 0.01% (v/v) | 5 mL |
| Total | n/a | 50 mL |
Prepare before use.
5. 3% sucrose (500 mL)
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Sucrose | 3% (w/v) | 30 g |
| dH2O | n/a | 500 mL |
| Total | n/a | 500 mL |
Filter the solution using a sterile bottle top filter inside a fume hood. Store at RT.
6. Extraction buffer (50 mL)
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| MeOH | 80% (v/v) | 40 mL |
| Formic acid | 3% (v/v) | 1.5 mL |
| dH2O | n/a | 8.5 mL |
| Total | n/a | 50 mL |
Store at RT.
7. HPLC running buffer A (1 L)
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Formic acid | 0.1% (v/v) | 1 mL |
| dH2O | n/a | 999 mL |
| Total | n/a | 1,000 mL |
Store at RT.
8. HPLC running buffer B (1 L)
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Formic acid | 0.1% (v/v) | 1 mL |
| Acetonitrile | n/a | 999 mL |
| Total | n/a | 1,000 mL |
Store at RT.
Laboratory supplies
1. Pipette tips 20–200 μL and 100–1,000 μL (FisherScientific, catalog numbers: 94052320, 94052410)
2. Falcon tubes 15 and 50 mL (FALCON Corning Products, catalog numbers: 352196, 352098)
3. 1.7 mL graduated microcentrifuge tubes (VWR, catalog number: 10025-716)
4. 60 × 15 mm Petri dish (VWR, catalog number: 77589-162)
5. 500 mL bottle top filter (FisherScientific, catalog number: FB12566510)
6. Parafilm (Bemis, catalog number: PM-992)
7. Weighing paper (VWR, catalog number: 12578-121)
8. HPLC vial (FisherScientific, catalog number: C4000-12)
9. HPLC vial insert (MUHWA Scientific, catalog number: X003K4XYWZ)
10. HPLC screw cup (Alwsci technologies, catalog number: C0002896)
11. Symmetry C18 column, 100 Å, 3.5 µm, 4.6 mm × 150 mm (Waters Corporation, catalog number: WAT200632)
12. Semi-micro cuvette (MiliporeSigma, catalog number: BR759015)
Equipment
1. Pipetteman pipettes (Gilson, catalog number: FD10006)
2. Graduated cylinder (PYREX, catalog number: 3042)
3. CorningTM PYREXTM reusable media storage bottles, Pyrex (FisherScientific, catalog number: 10-462-711)
4. Philips 30W 36in T12 cool white fluorescent tube (Bulbs.com, SKU: 272427)
5. Digital orbital shaker (ONiLAB, SKU: SK-O180-S)
6. Spectrometer (LI-COR Biosciences, model: LI-180)
7. Stainless steel forceps (MiliporeSigma, catalog number: Z168793)
8. Drawing pin (Amazon, catalog number: DH05101901)
9. -80 °C ultra-Low freezer (PHCbi, catalog number: MDF-DU502VH)
10. FreeZone Benchtop Freeze Dryer (Labconcom, catalog number: 700201000)
11. Vortex (Scientific Industries, catalog number: SI-0236)
12. Microcentrifuge (VWR, catalog number: VWRI521-2656)
13. HPLC system (Agilent Technologies, model: Agilent 1260)
14. Euromex NexiusZoom Series high-precision zoom binocular stereomicroscope (AmScope, catalog number: NZ1902-P)
15. Camera (AmScope, catalog number: MU1803)
16. Fine balance (FisherScientific, catalog number: 02-035SD)
17. Branson UltrasonicsTM CPX8800H ultrasonic cleaning bath (FisherScientific, catalog number: 15-336-124)
18. 0.22 mm filter (VWR, catalog number: 76478-992)
19. Digital timer (VWR, catalog number: 609-0204)
20. UV-1600PC scanning spectrophotometer (VWR, catalog number: 10037-436)
Software and datasets
1. AmScope version 4.12.24446.20240114 (Irvine, California)
2. Agilent OpenLab Control Panel, version 3.5.0 (Santa Clara, California)
3. Agilent OpenLab Acquisition, Build 2.6.0.691 (Santa Clara, California)
4. Agilent OpenLab Data Analysis, Build 2.206.0.780 (Santa Clara, California)
5. GraphPad Prism 10.1.1 (San Diego, California)
Procedure
文章信息
稿件历史记录
提交日期: Dec 17, 2025
接收日期: Feb 1, 2026
在线发布日期: Feb 12, 2026
出版日期: Mar 5, 2026
版权信息
© 2026 The Author(s); This is an open access article under the CC BY-NC license (https://creativecommons.org/licenses/by-nc/4.0/).
如何引用
He, Y., Iha, S., Grotewold, E. and Jiang, N. (2026). Simple Induction and Detection of Anthocyanins in Arabidopsis thaliana: A Tool for Mutant Screening and Metabolic Analysis. Bio-protocol 16(5): e5628. DOI: 10.21769/BioProtoc.5628.
分类
植物科学 > 植物生物化学 > 代谢物
生物化学 > 其它化合物 > 小分子
您对这篇实验方法有问题吗?
在此处发布您的问题,我们将邀请本文作者来回答。同时,我们会将您的问题发布到Bio-protocol Exchange,以便寻求社区成员的帮助。
Share
Bluesky
X
Copy link