- EN - English
- CN - 中文
Dynamic Mapping of RNA-Binding Proteins During Bacillus subtilis Sporulation Using Orthogonal Organic Phase Separation
基于正交有机相分离技术动态解析枯草芽孢杆菌孢子形成过程中 RNA 结合蛋白图谱
发布: 2026年03月05日第16卷第5期 DOI: 10.21769/BioProtoc.5622 浏览次数: 116
评审: Shengze YaoJing LiAnu Thomas
Abstract
RNA-binding proteins (RBPs) have pleiotropic roles in modulating the physiology of both eukaryotic and prokaryotic cells, enabling them to adapt to environmental variations. The importance of RBPs has led to the development of a variety of methods aiming to identify them. However, most of these approaches have primarily been implemented and optimized in eukaryotic systems. To both uncover novel RBPs involved in Bacillus subtilis sporulation and capture their RNA-binding ability dynamically, we adapted the orthogonal organic phase separation technique (OOPS), which had previously been used in Escherichia coli to reveal its RNA-binding proteome (RBPome). We optimized the UV cross-linking process used to stabilize RNA–protein interactions in vivo and the bacterial lysis process to overcome the robust cell wall of Gram-positive sporulating cells. RNA–protein complexes are then recovered after phase separation steps using guanidinium thiocyanate–phenol–chloroform, and RNA-associated proteins are identified and label-free-quantified by liquid chromatography–mass spectrometry. Collecting samples at various time points during sporulation further enables tracking the dynamics of the RBPome. In addition to being applicable to bacteria and requiring minimal starting material, this method has provided a comprehensive map of the RBPome during sporulation, refining the roles of known factors and revealing new players.
Key features
• The high-throughput method OOPS, developed in [1], was successfully applied to both sporulating and vegetative cells of a Gram-positive bacterium to specifically purify the RBPome.
• OOPS allowed tracking of RBPome remodeling dynamics (both RBP abundance and RNA binding ability) across different stages of B. subtilis sporulation.
• OOPS enabled the identification of novel RBPs in the context of sporulation, revealing potential new players in RNA-mediated regulation.
Keywords: RNA binding proteins (RNA 结合蛋白)Graphical overview
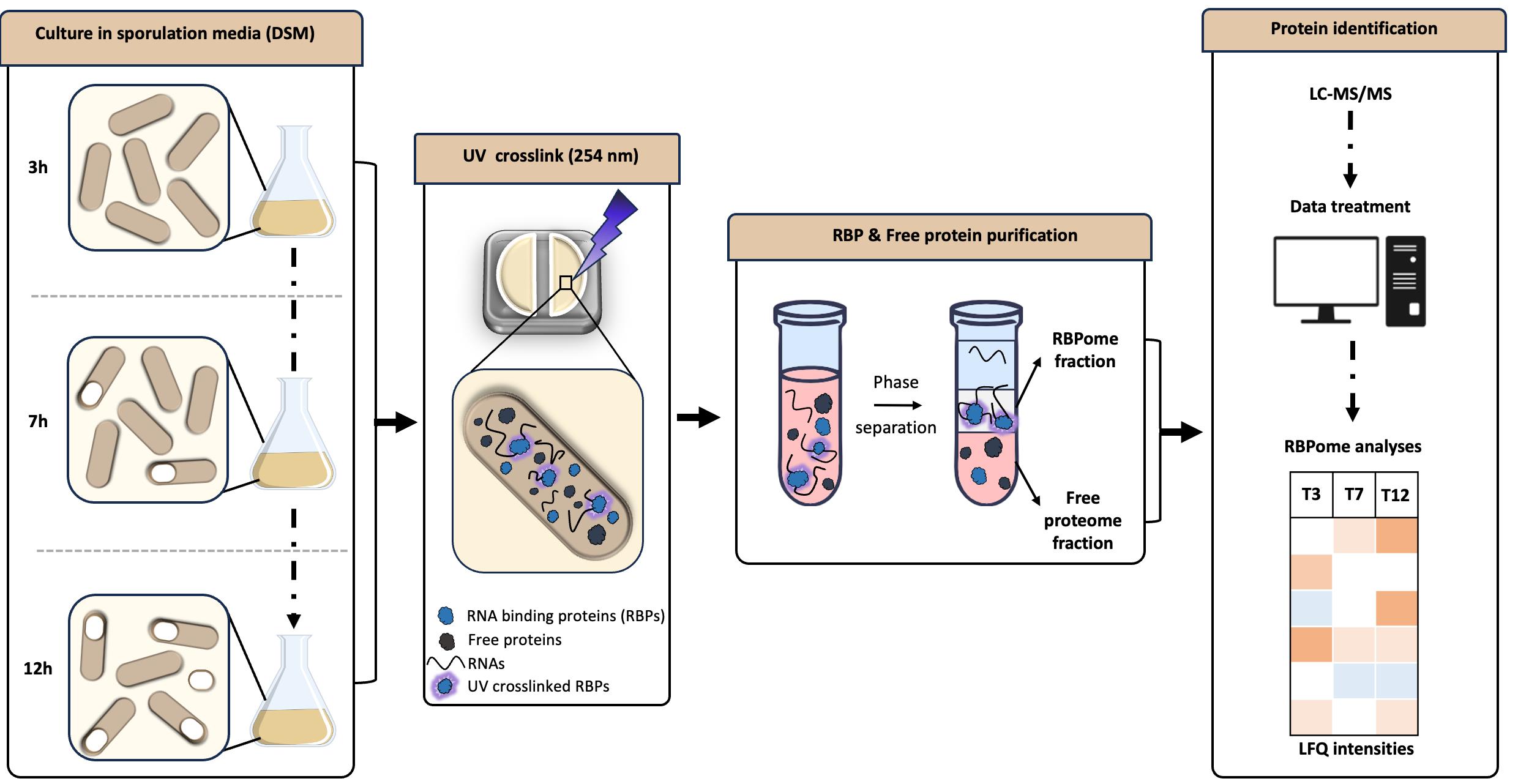
Background
RNA-binding proteins (RBPs) have emerged as key post-transcriptional regulators that help bacteria to adjust gene expression, especially in the face of rapid environmental changes. Based on the global transcriptome study of Nicolas et al. [2] in Bacillus subtilis, we observed that sporulation is the condition associated with the highest number of expressed putative small regulatory RNAs (sRNAs). While this suggests extensive post-transcriptional regulation during sporulation, its impact on the process remains largely unexplored. Given the diversity of both RNA–RBP interactions involving multiple classes of RNAs and RBP functions, a global approach centered on RNA–protein interactions was required to understand the complexity of these regulatory networks.
Sporulation is a highly dynamic process governed by sequential regulatory cascades known to act at transcriptional, translational, and post-translational levels. Therefore, to capture this temporal complexity, we adopted a dynamic strategy that would not only identify RBPs but also monitor how RNA–protein interactions are modulated over time. To achieve this, in our recent work on B. subtilis, we applied a global proteome-wide approach that would allow tracking the remodeling of the RBPome during the sporulation process and identify new candidate RBPs for further functional characterization.
Global mapping of RNA–protein interactions in bacteria remains technically challenging. Most RBPome methods were initially developed for eukaryotes and rely on poly(A) selection or large amounts of starting material, making them poorly suited to non-polyadenylated bacterial transcriptomes and to conditions in which biomass is limiting. Orthogonal organic phase separation (OOPS) overcomes several of these constraints by coupling UV cross-linking with TRIzol-based phase separation to enrich RNA–protein complexes directly from cell lysates in a sequence-independent manner [3,4]. Compared with related phase-extraction methods, OOPS required less input (for human cells, ~3 × 106 cells for OOPS vs. ~5 × 106 for PTex and ~8 × 107 for XRNAX) [3,4], which is a major advantage for time-resolved experiments. Although no systematic head-to-head comparison is available, published datasets suggest that OOPS achieves a depth of coverage comparable to alternative strategies; for instance, PTex recovered 384 enriched RBPs in Staphylococcus aureus, a number similar to the 364 E. coli RBPs identified using OOPS [5]. Additionally, the OOPS protocol is technically accessible, cost-effective, and rapid to implement, requiring no highly specialized expertise beyond standard proteomics and basic RNA handling. In our study, we used label-free quantification, which is less precise than isotope-based approaches such as SILAC but provides sufficient quantitative resolution to capture global RBPome remodeling and relative changes in RNA binding over the course of sporulation, while substantially reducing cost and experimental complexity.
OOPS was therefore ideally suited to our objective of obtaining an overview of the RBPome during sporulation and identifying new post-transcriptional regulators of this process. We applied OOPS to cultures sampled at different stages of sporulation—from vegetative cells to the appearance of the first mature spores—enabling dynamic tracking of the RBPome modulation during the process. Prior to our study, bacterial applications of OOPS had focused only on Gram-negative E. coli and had not yet been extended to a Gram-positive bacterium [3]. Consequently, we also had to adapt the UV cross-linking and cell lysis procedures to B. subtilis. Indeed, the cell wall of Gram-positive bacteria is composed of a thick layer of peptidoglycan and is more resistant than that of Gram-negative bacteria. B. subtilis forms prespores and highly resistant spores that require specific lysis conditions and show reduced sensitivity to UV treatment. This adapted protocol, based largely on the original OOPS workflow [4], includes targeted methodological adjustments that are in line with our study model. This approach allowed us to detect not only proteins that have already been described as RBP but also to provide dynamic monitoring of their binding to RNA during sporulation. Moreover, we identified a new candidate RBP that is particularly enriched during this process. Together, these results broaden our understanding of the proteins that may contribute to post-transcriptional regulation during sporulation [1].
Materials and reagents
Biological materials
1. B. subtilis WT168 (laboratory stock)
Reagents
1. BactoTM tryptone (Thermo Fisher Scientific, catalog number: 211705); store at room temperature (RT) in a dry place
2. BactoTM yeast extract (Thermo Fisher Scientific, catalog number: 212750); store at RT in a dry place
3. NaCl (Sigma-Aldrich, catalog number: S9888); store at RT in a dry place
4. Nutrient broth No. 4 (DSM) (Fluka Analytical, catalog number: 03856); store at RT in a dry place
5. KCl (Thermo Scientific, catalog number: 7447-40-7); store at RT in a dry place
6. MgSO4 (Sigma-Aldrich, catalog number: M1880-500G); store at RT in a dry place
7. Phosphate buffer saline (PBS) tablets, pH 7.4 (Euromedex, catalog number: EU1-2052-100); store at RT in a dry place
8. RNaseZapTM RNase decontamination solution (Invitrogen, catalog number: AM9782); store at RT
9. Lysing matrix B bulk (MP Biomedicals, catalog number: 6540425); store at RT in a dry place
10. TRIzol (Thermo Fisher Scientific, catalog number: 15596018), store at 4°C
Caution: This chemical is corrosive, irritating, and toxic.
11. Chloroform (Carlo Erba, catalog number: 438581); store at RT
Caution: This chemical is toxic.
12. Propan-2-ol (Carlo Erba, catalog number: 415154); store at RT
Caution: This chemical is flammable and irritating.
13. Phenol:chloroform:isoamyl alcohol (25:24:1, v/v) (PanReac AppliChem, catalog number: A0944); store at 4 °C
Caution: This chemical is corrosive, irritating, and toxic.
14. Ethanol absolute anhydrous (Carlo Erba, catalog number: 4146012); store at RT
Caution: This chemical is flammable.
15. Methanol (Carlo Erba, catalog number: 414819); store at RT
Caution: This chemical is flammable and toxic.
16. Proteinase K, recombinant PCR grade (Roche, catalog number: 03115887001); store at -20 °C
17. Sodium acetate (3 M, pH 5.5) (Thermo Fisher Scientific, catalog number: AM9740); store at 4 °C
18. EDTA (Sigma-Aldrich, catalog number: EDS); store at RT
Caution: This chemical is irritating and toxic.
19. RNace-It (RNase A/T1) cocktail (Agilent, catalog number: 400720); store at -20 °C
20. MgCl2 (Sigma-Aldrich, catalog number: M8266); store at RT
21. Tris-base (Euromedex, catalog number: 26-128-3094-B); store at RT
22. Sodium dodecyl sulfate (SDS) (Sigma-Aldrich, catalog number: M8266); store at RT
23. Triethylammonium bicarbonate (TEAB) buffer (1 M, pH 8.5) (Sigma-Aldrich, catalog number: T708); store at 4 °C
24. UltraPure water from PURELAB flex system (ELGA); store at RT
25. Imperial Blue staining solution (Thermo Fisher Scientific, catalog number: 24615); store at RT protected from light
26. Sequencing-grade modified trypsin (Promega, catalog number: V5111); store at -20 °C
27. Ammonium bicarbonate (ABC) (Sigma/Merck, catalog number: 09830); store at RT
28. Iodoacetamide (IAA) (Sigma/Merck, catalog number: I1149); store at -20 °C
29. Acetonitrile LCMS (ACN) (Sigma/Merck, catalog number: 1.00029); store at RT
30. DL-Dithiothreitol (DTT) (Sigma/Merck, catalog number: D9163); store at -20 °C
31. Color protein standard (New England Biolabs, catalog number: P7719S)
32. NuPAGETM 4%–12% Bis-Tris gel (Invitrogen, catalog numbers: NP0321BOX and NP0322BOX)
33. NuPAGETM MOPS SDS running buffer 20× (Invitrogen, catalog number: NP001)
Solutions
1. LB media (see Recipes)
2. DSM stock solution (see Recipes)
3. DSM working solution (see Recipes)
4. TN150 solution (see Recipes)
5. Lysis solution (see Recipes)
6. 10 mM DTT (see Recipes)
7. 55 mM IAA (see Recipes)
8. 20× Trypsin/Lys-C (see Recipes)
9. 1× Trypsin/Lys-C mix (see Recipes)
Recipes
1. LB media
| Reagent | Final concentration | Volume |
|---|---|---|
| Bacto tryptone | 10 g/L | 10 g |
| Yeast extract | 5 g/L | 5 g |
| NaCl | 5 g/L | 5 g |
| Demineralized water | n/a | 1 L |
| Total | n/a | 1 L |
2. DSM stock solution
| Reagent | Final concentration | Volume |
|---|---|---|
| Difco nutrient broth | 8 g/L | 8 g |
| KCl | 1 g/L | 1 g |
| MgSO4·7H2O | 0.25 g/L | 0.25 g |
| Demineralized water | n/a | 1 L |
| Total | n/a | 1 L |
3. DSM working solution
| Reagent | Final concentration | Volume |
|---|---|---|
| DSM stock solution | n/a | 10 mL |
| 1 M Ca(NO3)2 | 1 mM | 10 μL |
| 10 mM MnCl2 | 10 μM | 10 μL |
| 83 mM FAC (ferric ammonium citrate) | 80 μM | 10 μL |
| 1 mM FeSO4 | 200 nM | 2 μL |
| Total | n/a | 10 mL |
All elements must be added on the day of the culture.
4. TN150 solution
| Reagent | Final concentration | Volume |
|---|---|---|
| Tris-HCl pH 7.8 (1.0 M) | 50 mM | 250 μL |
| NaCl (5.0 M) | 150 mM | 150 μL |
| NP-40/IGEPAL CA-630 (10%) | 0.1% | 50 μL |
| RNase-free H2O | n/a | 4.55 mL |
| Total | n/a | 5 mL |
TN150 can be stored at 4 °C for 1–3 months.
5. Lysis solution
| Reagent | Final concentration | Volume |
|---|---|---|
| TN150 | n/a | 1.72 mL |
| Lysozyme (100 mg/mL) | 10 mg/mL | 200 μL |
| DNase RQ1 (1 U/μL) | 0.035 U/μL | 80 μL |
| Total | n/a | 2 mL |
Prepare freshly prior to use and avoid storage.
6. 10 mM DTT
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| 100 mM ammonium bicarbonate (ABC) | 100 mM | 1.5 mL |
| DTT | 1.54 g/L | 2.3 mg |
| Total | n/a | 1.5 mL |
Aliquot and store at -20 °C for 3–6 months.
7. 55 mM IAA
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| 100 mM ABC | 100 mM | 1.5 mL |
| IAA | 10.17 g/L | 15.3 mg |
| Total | n/a | 1.5 mL |
Aliquot and store at -20 °C for 3–6 months protected from light.
8. 20× Trypsin/Lys-C
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Manufacturer resuspension buffer | n/a | 100 μL |
| Trypsin/Lys-C (lyophilized) | 200 ng/μL | 20 μg (lyophilized) |
| Total | n/a | 100 μL |
Aliquot and store at -20 °C for 6–12 months.
9. 1× Trypsin/Lys-C mix
| Reagent | Final concentration | Volume |
|---|---|---|
| 20× Trypsin/Lys-C | 10 ng/μL | 10 μL |
| 25 mM ABC | 25 mM | 185 μL |
| 1% ProteaseMax | 0.025% | 5 μL |
| Total | n/a | 200 μL |
Prepare freshly; avoid storage.
Laboratory supplies
1. 1 L culture flasks
2. INFORS HT Multitron shaking incubator
3. Sarstedt tubes 15 mL (Sarstedt, catalog number: 62.554.502) and 50 mL (Sarstedt, catalog number: 62.547.254)
4. Nylon filter membranes, 0.45 μm pore size, 90 mm (Merck Millipore, catalog number: HNWP09000)
5. Microfuge tubes 1.5 mL (Sarstedt, catalog number: 72.690.001)
6. Disposable gloves
7. Paper towels
8. Micropipettes (2 μL, 20 μL, 200 μL, 1 mL)
9. C18 trap column (0.3 × 5 mm) PepMap Neo C18 5 μm, 300 μm × 5 mm 17400 (Thermo Scientific, catalog number: 174500)
10. EASY-SprayTM PepMapTM Neo capillary column (75 μm × 50 cm or 75 μm × 15 cm, 2 μm, 100 Å) (Thermo Scientific, catalog numbers: ES75500PN or ES75750PN)
11. Scalpels (Swann Morton, catalog number: 0501)
12. Protein Lo-Bind tubes (Eppendorf, catalog number: 0030108116)
Equipment
1. Hybridization incubator (UVP, model: HL-2000)
2. DWK Life DURANTM filter funnel head, with PP funnel and two FKM seals (Fisher Scientific, catalog number: 10202321)
3. DWK Life Sciences DURANTM filter disc, fused glass rim (Fisher Scientific, catalog number: 10179560)
4. Thermo shaker (Thermal Shake lite) (VWR, catalog number: 460−0249)
5. FastPrep®-24 5G bead beating grinder and lysis system (MP Biomedicals)
6. Table-top vortex
7. Fume extraction hood
8. NanoDropTM One/OneC Microvolume UV-Visible Spectrophotometer (Thermo ScientificTM)
9. InvitrogenTM mini gel tank and transfer module set (Invitrogen, catalog number: 15384604)
10. S700 ChillBlockTM tube racks for MCT (Simport Scientific, catalog numbers: S700-16 and S700-14)
11. Centrifuge (Eppendorf, model: 5810/5810 R)
12. Centrifuge paillasse (Eppendorf)
13. SavantTM SpeedVacTM DNA 130 Integrated Vacuum Concentration System (Thermo Fisher)
14. Q-Exactive Plus (Orbitrap, Thermo Fisher)
Software and datasets
1. MaxQuant (v.2.5.0.0; downloaded from https://maxquant.net/maxquant/, Cox lab, Max Planck Institute of Biochemistry, Martinsried, Germany)
2. Perseus software (v 1.6.15)
3. Microsoft Excel (Microsoft 365)
Procedure
文章信息
稿件历史记录
提交日期: Dec 17, 2025
接收日期: Jan 27, 2026
在线发布日期: Feb 9, 2026
出版日期: Mar 5, 2026
版权信息
© 2026 The Author(s); This is an open access article under the CC BY-NC license (https://creativecommons.org/licenses/by-nc/4.0/).
如何引用
Kaboré, T. and Delan-Forino, C. (2026). Dynamic Mapping of RNA-Binding Proteins During Bacillus subtilis Sporulation Using Orthogonal Organic Phase Separation. Bio-protocol 16(5): e5622. DOI: 10.21769/BioProtoc.5622.
分类
微生物学 > 微生物蛋白质组学
分子生物学 > 蛋白质
您对这篇实验方法有问题吗?
在此处发布您的问题,我们将邀请本文作者来回答。同时,我们会将您的问题发布到Bio-protocol Exchange,以便寻求社区成员的帮助。
Share
Bluesky
X
Copy link